+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1noy | ||||||
|---|---|---|---|---|---|---|---|
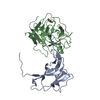
| Title | DNA POLYMERASE (E.C.2.7.7.7)/DNA COMPLEX | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSFERASE/DNA / EXONUCLEASE / DNA-BINDING / COMPLEX (NUCLEOTIDYLTRANSFERASE-DNA) / TRANSFERASE-DNA complex | ||||||
| Function / homology |  Function and homology information Function and homology informationbidirectional double-stranded viral DNA replication / Hydrolases; Acting on ester bonds; Exodeoxyribonucleases producing 5'-phosphomonoesters / 3'-5' exonuclease activity / DNA-templated DNA replication / DNA-directed DNA polymerase / DNA-directed DNA polymerase activity / nucleotide binding / DNA binding / metal ion binding Similarity search - Function | ||||||
| Biological species |  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.2 Å X-RAY DIFFRACTION / Resolution: 2.2 Å | ||||||
 Authors Authors | Wang, J. / Yu, P. / Lin, T.C. / Konigsberg, W.H. / Steitz, T.A. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1996 Journal: Biochemistry / Year: 1996Title: Crystal structures of an NH2-terminal fragment of T4 DNA polymerase and its complexes with single-stranded DNA and with divalent metal ions. Authors: Wang, J. / Yu, P. / Lin, T.C. / Konigsberg, W.H. / Steitz, T.A. #1:  Journal: J.Biol.Chem. / Year: 1994 Journal: J.Biol.Chem. / Year: 1994Title: Isolation, Characterization, and Kinetic Properties of Truncated Forms of T4 DNA Polymerase that Exhibit 3'-5' Exonuclease Activity Authors: Lin, T.C. / Karam, G. / Konigsberg, W.H. #2:  Journal: J.Biol.Chem. / Year: 1988 Journal: J.Biol.Chem. / Year: 1988Title: Primary Structure of T4 DNA Polymerase. Evolutionary Relatedness to Eucaryotic and Other Procaryotic DNA Polymerases Authors: Spicer, E.K. / Rush, J. / Fung, C. / Reha-Krantz, L.J. / Karam, J.D. / Konigsberg, W.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1noy.cif.gz 1noy.cif.gz | 160.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1noy.ent.gz pdb1noy.ent.gz | 126.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1noy.json.gz 1noy.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/no/1noy https://data.pdbj.org/pub/pdb/validation_reports/no/1noy ftp://data.pdbj.org/pub/pdb/validation_reports/no/1noy ftp://data.pdbj.org/pub/pdb/validation_reports/no/1noy | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 867.621 Da / Num. of mol.: 1 / Source method: obtained synthetically | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| #2: Protein | Mass: 45315.727 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Enterobacteria phage T4 (virus) / Genus: T4-like viruses / Species: Enterobacteria phage T4 sensu lato / Production host: Enterobacteria phage T4 (virus) / Genus: T4-like viruses / Species: Enterobacteria phage T4 sensu lato / Production host:  #3: Chemical | ChemComp-ZN / | #4: Chemical | ChemComp-MN / | #5: Water | ChemComp-HOH / | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.41 Å3/Da / Density % sol: 50 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal | *PLUS Density % sol: 50 % | ||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 7 / Method: unknown | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 298 K |
|---|---|
| Detector | Type: SDMS / Detector: AREA DETECTOR / Date: Oct 7, 1993 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | % possible obs: 100 % |
| Reflection | *PLUS Highest resolution: 2.2 Å / Rmerge(I) obs: 0.096 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.2→6 Å / σ(F): 2 Details: THIS STRUCTURE WAS DETERMINED AT ROOM TEMPERATURE (298K). THE DIVALENT AND P(DT)3 WERE DETERMINED FROM TWO SEPARATE EXPERIMENTS AT LOWER RESOLUTIONS, CONCATENATED TOGETHER IN THIS ENTRY. ASP ...Details: THIS STRUCTURE WAS DETERMINED AT ROOM TEMPERATURE (298K). THE DIVALENT AND P(DT)3 WERE DETERMINED FROM TWO SEPARATE EXPERIMENTS AT LOWER RESOLUTIONS, CONCATENATED TOGETHER IN THIS ENTRY. ASP 1, GLU 2, ILE 211 AND FOUR OTHER RESIDUES WERE IDENTIFIED FROM ELECTRON DENSITY MAPS, BUT THE AUTHORS HAVE NOT DONE DNA SEQUENCING TO CONFIRM POSSIBLE SPONTANEOUSLY MUTATIONS. ILE 211 WAS REPLACED BY THR 211 IN THIS ENTRY WITHOUT ADDITIONAL REFINEMENT BY REMOVING ATOM CD1.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.2 Å / Lowest resolution: 6 Å / σ(F): 2 / Rfactor obs: 0.222 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 39.8 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj







