+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1mk0 | ||||||
|---|---|---|---|---|---|---|---|
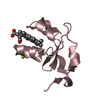
| Title | catalytic domain of intron endonuclease I-TevI, E75A mutant | ||||||
 Components Components | Intron-associated endonuclease 1 | ||||||
 Keywords Keywords | HYDROLASE / alpha/beta fold / CATALYTIC DOMAIN / DNA-BINDING SURFACE | ||||||
| Function / homology |  Function and homology information Function and homology informationdouble-stranded DNA endonuclease activity / intron homing / DNA-binding transcription repressor activity / transcription repressor complex / endonuclease activity / sequence-specific DNA binding / Hydrolases; Acting on ester bonds / negative regulation of DNA-templated transcription / DNA binding / zinc ion binding Similarity search - Function | ||||||
| Biological species |  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.6 Å MOLECULAR REPLACEMENT / Resolution: 1.6 Å | ||||||
 Authors Authors | Van Roey, P. / Meehan, L. / Kowalski, J.C. / Belfort, M. / Derbyshire, V. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2002 Journal: Nat.Struct.Biol. / Year: 2002Title: Catalytic domain structure and hypothesis for function of GIY-YIG intron endonuclease I-TevI. Authors: Van Roey, P. / Meehan, L. / Kowalski, J.C. / Belfort, M. / Derbyshire, V. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1mk0.cif.gz 1mk0.cif.gz | 37.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1mk0.ent.gz pdb1mk0.ent.gz | 24.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1mk0.json.gz 1mk0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mk/1mk0 https://data.pdbj.org/pub/pdb/validation_reports/mk/1mk0 ftp://data.pdbj.org/pub/pdb/validation_reports/mk/1mk0 ftp://data.pdbj.org/pub/pdb/validation_reports/mk/1mk0 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1ln0SC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 11406.057 Da / Num. of mol.: 1 / Fragment: catalytic domain (residues 1 to 97) / Mutation: E75A Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Enterobacteria phage T4 (virus) / Genus: T4-like viruses / Species: Enterobacteria phage T4 sensu lato / Production host: Enterobacteria phage T4 (virus) / Genus: T4-like viruses / Species: Enterobacteria phage T4 sensu lato / Production host:  References: UniProt: P13299, Hydrolases; Acting on ester bonds |
|---|---|
| #2: Chemical | ChemComp-CIT / |
| #3: Chemical | ChemComp-BME / |
| #4: Water | ChemComp-HOH / |
| Has protein modification | N |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.39 Å3/Da / Density % sol: 48.48 % | |||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 283 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: 1.6 M sodium citrate, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 283K | |||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 7.5 | |||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X12C / Wavelength: 1 Å / Beamline: X12C / Wavelength: 1 Å |
| Detector | Type: BRANDEIS - B1.2 / Detector: CCD / Date: Jul 11, 2002 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→50 Å / Num. all: 15342 / Num. obs: 15342 / % possible obs: 99.7 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -3 / Redundancy: 9 % / Biso Wilson estimate: 16.7 Å2 / Rmerge(I) obs: 0.034 / Net I/σ(I): 84 |
| Reflection shell | Resolution: 1.6→1.65 Å / Redundancy: 6.5 % / Rmerge(I) obs: 0.092 / Mean I/σ(I) obs: 12 / % possible all: 98.3 |
| Reflection | *PLUS Lowest resolution: 50 Å / Rmerge(I) obs: 0.034 |
| Reflection shell | *PLUS % possible obs: 98.3 % / Rmerge(I) obs: 0.09 / Mean I/σ(I) obs: 23.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: pdb entry 1LN0 Resolution: 1.6→36.55 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 1085373.69 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 41.2046 Å2 / ksol: 0.393289 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 16.6 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error free: 0.21 Å / Luzzati sigma a free: 0.11 Å | ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→36.55 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.6→1.7 Å / Rfactor Rfree error: 0.017 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.216 / Rfactor Rwork: 0.197 | ||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.262 / Rfactor Rwork: 0.229 |
 Movie
Movie Controller
Controller












 PDBj
PDBj







