[English] 日本語
 Yorodumi
Yorodumi- PDB-1jlp: Solution Structure of the Noncompetitive Skeletal Muscle Nicotini... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1jlp | ||||||
|---|---|---|---|---|---|---|---|
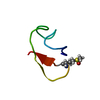
| Title | Solution Structure of the Noncompetitive Skeletal Muscle Nicotinic Acetylcholine Receptor Antagonist Psi-conotoxin PIIIF | ||||||
 Components Components | PSI-CONOTOXIN PIIIF | ||||||
 Keywords Keywords | TOXIN / MULTIPLE DISULFIDE BONDS / AMIDATED C-TERMINUS | ||||||
| Function / homology | host cell postsynaptic membrane / toxin activity / extracellular region / Psi-conotoxin PIIIF Function and homology information Function and homology information | ||||||
| Method | SOLUTION NMR / distance geometry molecular dynamics relaxation matrix distance geometry simulated annealing | ||||||
 Authors Authors | Van Wagoner, R.M. / Ireland, C.M. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2003 Journal: Biochemistry / Year: 2003Title: Characterization and Three-Dimensional Structure Determination of psi-Conotoxin Piiif, a Novel Noncompetitive Antagonist of Nicotinic Acetylcholine Receptors Authors: Van Wagoner, R.M. / Ireland, C.M. #1:  Journal: Biochemistry / Year: 1998 Journal: Biochemistry / Year: 1998Title: Three-dimensional Solution Structure of Conotoxin psi-PIIIE, an Acetylcholine Gated Ion Channel Antagonist Authors: Mitchell, S.S. / Shon, K.J. / Foster, M.P. / Davis, D.R. / Olivera, B.M. / Ireland, C.M. #2:  Journal: Biochemistry / Year: 1997 Journal: Biochemistry / Year: 1997Title: A Noncompetitive Peptide Inhibitor of the Nicotinic Acetylcholine Receptor from Conus purpurascens Venom Authors: Shon, K.J. / Grilley, M. / Jacobsen, R. / Cartier, G.E. / Hopkins, C. / Gray, W.R. / Watkins, M. / Hillyard, D.R. / Rivier, J. / Torres, J. / Yoshikami, D. / Olivera, B.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1jlp.cif.gz 1jlp.cif.gz | 106.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1jlp.ent.gz pdb1jlp.ent.gz | 86.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1jlp.json.gz 1jlp.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1jlp_validation.pdf.gz 1jlp_validation.pdf.gz | 361 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1jlp_full_validation.pdf.gz 1jlp_full_validation.pdf.gz | 448.4 KB | Display | |
| Data in XML |  1jlp_validation.xml.gz 1jlp_validation.xml.gz | 8.6 KB | Display | |
| Data in CIF |  1jlp_validation.cif.gz 1jlp_validation.cif.gz | 13.4 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jl/1jlp https://data.pdbj.org/pub/pdb/validation_reports/jl/1jlp ftp://data.pdbj.org/pub/pdb/validation_reports/jl/1jlp ftp://data.pdbj.org/pub/pdb/validation_reports/jl/1jlp | HTTPS FTP |
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 2674.157 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: Solid phase chemical synthesis was used. One sample contained only natural isotopic abundance residues. Another peptide incorporated 3-13C-labeled cysteine at position 4. The third peptide ...Details: Solid phase chemical synthesis was used. One sample contained only natural isotopic abundance residues. Another peptide incorporated 3-13C-labeled cysteine at position 4. The third peptide incorporated 3-13C-labeled cysteine at positions 5, 10, 16, 21, and 22. The sequence for all peptides is the same as the native peptide FROM CONUS PURPURASCENS (PURPLE CONE). References: UniProt: P60245 |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||||||||||
| NMR details | Text: This structure was determined using isotopic filtration and isotope-selected 1H observation in addition to standard homonuclear techniques. |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions |
| ||||||||||||||||||
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | |||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: distance geometry molecular dynamics relaxation matrix distance geometry simulated annealing Software ordinal: 1 Details: These structures are based on 380 NOE-derived distance restraints, 7 restraints on phi derived from J-coupling, and 7 restraints on chi1 determined from J-coupling and NOE volumes. | ||||||||||||||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: back calculated data agree with experimental NOESY spectrum, structures with the least restraint violations, structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 17 |
 Movie
Movie Controller
Controller










 PDBj
PDBj