[English] 日本語
 Yorodumi
Yorodumi- PDB-1iqd: Human Factor VIII C2 Domain complexed to human monoclonal BO2C11 Fab. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1iqd | ||||||
|---|---|---|---|---|---|---|---|
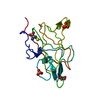
| Title | Human Factor VIII C2 Domain complexed to human monoclonal BO2C11 Fab. | ||||||
 Components Components |
| ||||||
 Keywords Keywords | IMMUNE SYSTEM/BLOOD CLOTTING / FACTOR VIII / C2 DOMAIN / ANTIBODY / BLOOD COAGULATION / INHIBITOR / BO2C11 / IMMUNE SYSTEM-BLOOD CLOTTING COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationDefective F8 accelerates dissociation of the A2 domain / Defective F8 binding to the cell membrane / Defective F8 secretion / Defective F8 sulfation at Y1699 / Gamma carboxylation, hypusinylation, hydroxylation, and arylsulfatase activation / Defective F8 binding to von Willebrand factor / blood coagulation, intrinsic pathway / Cargo concentration in the ER / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant ...Defective F8 accelerates dissociation of the A2 domain / Defective F8 binding to the cell membrane / Defective F8 secretion / Defective F8 sulfation at Y1699 / Gamma carboxylation, hypusinylation, hydroxylation, and arylsulfatase activation / Defective F8 binding to von Willebrand factor / blood coagulation, intrinsic pathway / Cargo concentration in the ER / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant / Defective F9 variant does not activate FX / COPII-coated ER to Golgi transport vesicle / COPII-mediated vesicle transport / Defective F8 cleavage by thrombin / : / : / endoplasmic reticulum-Golgi intermediate compartment membrane / platelet alpha granule lumen / acute-phase response / Golgi lumen / blood coagulation / Platelet degranulation / oxidoreductase activity / endoplasmic reticulum lumen / copper ion binding / : / extracellular region / plasma membrane Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Spiegel Jr., P.C. / Jacquemin, M. / Saint-Remy, J.M. / Stoddard, B.L. / Pratt, K.P. | ||||||
 Citation Citation |  Journal: Blood / Year: 2001 Journal: Blood / Year: 2001Title: Structure of a factor VIII C2 domain-immunoglobulin G4kappa Fab complex: identification of an inhibitory antibody epitope on the surface of factor VIII. Authors: Spiegel Jr., P.C. / Jacquemin, M. / Saint-Remy, J.M. / Stoddard, B.L. / Pratt, K.P. #1:  Journal: Nature / Year: 1999 Journal: Nature / Year: 1999Title: Structure of the C2 domain of human factor VIII at 1.5 A resolution Authors: Pratt, K.P. / Shen, B.W. / Takeshima, K. / Davie, E.W. / Fujikawa, K. / Stoddard, B.L. #2:  Journal: Blood / Year: 2000 Journal: Blood / Year: 2000Title: Hemophilic factor VIII C1- and C2-domain missense mutations and their modeling to the 1.5-angstrom human C2-domain crystal structure Authors: Liu, M.L. / Shen, B.W. / Nakaya, S. / Pratt, K.P. / Fujikawa, K. / Davie, E.W. / Stoddard, B.L. / Thompson, A.R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1iqd.cif.gz 1iqd.cif.gz | 133.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1iqd.ent.gz pdb1iqd.ent.gz | 101.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1iqd.json.gz 1iqd.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/iq/1iqd https://data.pdbj.org/pub/pdb/validation_reports/iq/1iqd ftp://data.pdbj.org/pub/pdb/validation_reports/iq/1iqd ftp://data.pdbj.org/pub/pdb/validation_reports/iq/1iqd | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Antibody | Mass: 22920.434 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Homo sapiens (human) / Strain: BO2C11 Homo sapiens (human) / Strain: BO2C11 |
|---|---|
| #2: Antibody | Mass: 22738.371 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Homo sapiens (human) / Strain: BO2C11 Homo sapiens (human) / Strain: BO2C11 |
| #3: Protein | Mass: 17744.326 Da / Num. of mol.: 1 / Fragment: C-TERMINAL DOMAIN / Mutation: S2296C Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Plasmid: pPIC9K / Production host: Homo sapiens (human) / Plasmid: pPIC9K / Production host:  Pichia pastoris (fungus) / Strain (production host): GS115 / References: UniProt: P00451 Pichia pastoris (fungus) / Strain (production host): GS115 / References: UniProt: P00451 |
| #4: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.11 Å3/Da / Density % sol: 41.62 % | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7 Details: PEG8000, sodium chloride, HEPES, pH 7.0, VAPOR DIFFUSION, HANGING DROP, temperature 298K | ||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7.5 / Method: vapor diffusion | ||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALS ALS  / Beamline: 5.0.2 / Wavelength: 1.1 Å / Beamline: 5.0.2 / Wavelength: 1.1 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: May 10, 2000 |
| Radiation | Monochromator: ALS 5.0.2 / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.1 Å / Relative weight: 1 |
| Reflection | Resolution: 2→50 Å / Num. all: 284482 / Num. obs: 284362 / % possible obs: 99.6 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Biso Wilson estimate: 10.9 Å2 |
| Reflection shell | Resolution: 2→2.03 Å / % possible all: 98.8 |
| Reflection | *PLUS Lowest resolution: 50 Å / Num. obs: 36867 / Redundancy: 7.7 % / Num. measured all: 284482 / Rmerge(I) obs: 0.061 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2→19.58 Å / Rfactor Rfree error: 0.004 / Data cutoff high absF: 225052.78 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 2→19.58 Å / Rfactor Rfree error: 0.004 / Data cutoff high absF: 225052.78 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 55.966 Å2 / ksol: 0.361134 e/Å3 | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24.5 Å2
| ||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→19.58 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.13 Å / Rfactor Rfree error: 0.012 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 0 / % reflection Rfree: 9.9 % / Rfactor obs: 0.203 | ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 24.5 Å2 | ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.271 / % reflection Rfree: 9.4 % / Rfactor Rwork: 0.213 |
 Movie
Movie Controller
Controller












 PDBj
PDBj











