+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1du0 | ||||||
|---|---|---|---|---|---|---|---|
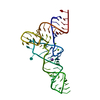
| Title | ENGRAILED HOMEODOMAIN Q50A VARIANT DNA COMPLEX | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSCRIPTION/DNA / homeodomain / DNA-binding protein / protein-DNA complex / TRANSCRIPTION-DNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationposterior compartment specification / analia development / anterior head segmentation / anterior/posterior lineage restriction, imaginal disc / genital disc development / genital disc anterior/posterior pattern formation / posterior head segmentation / trunk segmentation / spiracle morphogenesis, open tracheal system / wing disc anterior/posterior pattern formation ...posterior compartment specification / analia development / anterior head segmentation / anterior/posterior lineage restriction, imaginal disc / genital disc development / genital disc anterior/posterior pattern formation / posterior head segmentation / trunk segmentation / spiracle morphogenesis, open tracheal system / wing disc anterior/posterior pattern formation / wing disc morphogenesis / imaginal disc-derived wing vein specification / neuroblast fate determination / segment polarity determination / ventral midline development / compartment pattern specification / gonad development / axon guidance / RNA polymerase II transcription regulatory region sequence-specific DNA binding / DNA-binding transcription repressor activity, RNA polymerase II-specific / neuron differentiation / regulation of gene expression / sequence-specific DNA binding / negative regulation of neuron apoptotic process / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / negative regulation of gene expression / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / nucleus Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2 Å X-RAY DIFFRACTION / Resolution: 2 Å | ||||||
 Authors Authors | Grant, R.A. / Rould, M.A. / Klemm, J.D. / Pabo, C.O. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2000 Journal: Biochemistry / Year: 2000Title: Exploring the role of glutamine 50 in the homeodomain-DNA interface: crystal structure of engrailed (Gln50 --> ala) complex at 2.0 A. Authors: Grant, R.A. / Rould, M.A. / Klemm, J.D. / Pabo, C.O. #1:  Journal: J.Mol.Biol. / Year: 1998 Journal: J.Mol.Biol. / Year: 1998Title: Engrailed homeodomain-DNA complex at 2.2 Angstroms resolution: A detailed view of the interface and comparison with other engrailed structures Authors: Fraenkel, E. / Rould, M.A. / Chambers, K.A. / Pabo, C.O. #2:  Journal: Structure / Year: 1997 Journal: Structure / Year: 1997Title: Engrailed (gln50->lys) homeodomain-DNA complex at 1.9 Angstroms resolution: structural basis for enhanced affinity and altered specificity Authors: Tucker-Kellogg, L. / Rould, M.A. / Chambers, K.A. / Ades, S.E. / Sauer, R.T. #3:  Journal: Biochemistry / Year: 1994 Journal: Biochemistry / Year: 1994Title: Differential DNA-binding specificity of the engrailed homeodomain: the role of residue 50 Authors: Ades, S.E. / Sauer, R.T. #4:  Journal: Cell(Cambridge,Mass.) / Year: 1990 Journal: Cell(Cambridge,Mass.) / Year: 1990Title: Crystal structure of an engrailed homeodomain-DNA comples at 2.8 Angstroms resolution: a framework for understanding homeodomain-DNA interactions Authors: Kissinger, C.A. / Liu, B. / Martin-Blanco, E. / Kornberg, T.B. / Pabo, C.O. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1du0.cif.gz 1du0.cif.gz | 60.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1du0.ent.gz pdb1du0.ent.gz | 42.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1du0.json.gz 1du0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/du/1du0 https://data.pdbj.org/pub/pdb/validation_reports/du/1du0 ftp://data.pdbj.org/pub/pdb/validation_reports/du/1du0 ftp://data.pdbj.org/pub/pdb/validation_reports/du/1du0 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 6387.157 Da / Num. of mol.: 1 / Source method: obtained synthetically | ||
|---|---|---|---|
| #2: DNA chain | Mass: 6494.248 Da / Num. of mol.: 1 / Source method: obtained synthetically | ||
| #3: Protein | Mass: 6907.921 Da / Num. of mol.: 2 / Mutation: GLN50ALA Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.31 Å3/Da / Density % sol: 62.8 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 295 K / Method: vapor diffusion, hanging drop / pH: 7 Details: bis-Tris-propane, PEG 400, ammonium acetate, pH 7.0, VAPOR DIFFUSION, HANGING DROP, temperature 295K | ||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 128 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS IIC / Detector: IMAGE PLATE / Date: Jan 24, 1996 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2→20 Å / Num. all: 23204 / Num. obs: 23204 / % possible obs: 95.5 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 4.9 % / Biso Wilson estimate: 24.7 Å2 / Rmerge(I) obs: 0.039 |
| Reflection shell | Resolution: 2→2.07 Å / Redundancy: 3 % / Rmerge(I) obs: 0.351 / Num. unique all: 1697 / % possible all: 70.3 |
| Reflection | *PLUS Num. measured all: 115772 |
| Reflection shell | *PLUS % possible obs: 70.3 % / Mean I/σ(I) obs: 2.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2→20 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 10000000 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 39.7 Å2 | ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→20 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.13 Å / Rfactor Rfree error: 0.025 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.851 / Classification: refinement X-PLOR / Version: 3.851 / Classification: refinement | ||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.269 | ||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj







































