+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1bmp | ||||||
|---|---|---|---|---|---|---|---|
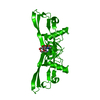
| Title | BONE MORPHOGENETIC PROTEIN-7 | ||||||
 Components Components | BONE MORPHOGENETIC PROTEIN-7 | ||||||
 Keywords Keywords | TRANSFORMING GROWTH FACTOR / MORPHOGEN / CYTOKINE / BONE / CARTILAGE / GLYCOPROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of prostatic bud formation / negative regulation of mesenchymal cell apoptotic process involved in nephron morphogenesis / negative regulation of glomerular mesangial cell proliferation / mesenchymal cell apoptotic process involved in nephron morphogenesis / positive regulation of cardiac neural crest cell migration involved in outflow tract morphogenesis / positive regulation of hyaluranon cable assembly / chorio-allantoic fusion / metanephric mesenchyme morphogenesis / nephrogenic mesenchyme morphogenesis / embryonic skeletal joint morphogenesis ...negative regulation of prostatic bud formation / negative regulation of mesenchymal cell apoptotic process involved in nephron morphogenesis / negative regulation of glomerular mesangial cell proliferation / mesenchymal cell apoptotic process involved in nephron morphogenesis / positive regulation of cardiac neural crest cell migration involved in outflow tract morphogenesis / positive regulation of hyaluranon cable assembly / chorio-allantoic fusion / metanephric mesenchyme morphogenesis / nephrogenic mesenchyme morphogenesis / embryonic skeletal joint morphogenesis / neural fold elevation formation / negative regulation of striated muscle cell apoptotic process / mesenchyme development / metanephric mesenchymal cell proliferation involved in metanephros development / positive regulation of epithelial cell differentiation / ameloblast differentiation / embryonic camera-type eye morphogenesis / allantois development / mesonephros development / monocyte aggregation / regulation of removal of superoxide radicals / pericardium morphogenesis / hindbrain development / mesenchymal cell differentiation / negative regulation of mitotic nuclear division / BMP receptor binding / endocardial cushion formation / pharyngeal system development / cellular response to BMP stimulus / branching involved in salivary gland morphogenesis / heart trabecula morphogenesis / embryonic pattern specification / metanephros development / cartilage development / negative regulation of non-canonical NF-kappaB signal transduction / embryonic limb morphogenesis / positive regulation of heterotypic cell-cell adhesion / response to vitamin D / positive regulation of dendrite development / ureteric bud development / cardiac muscle tissue development / Molecules associated with elastic fibres / regulation of branching involved in prostate gland morphogenesis / branching morphogenesis of an epithelial tube / cardiac septum morphogenesis / negative regulation of neuron differentiation / odontogenesis of dentin-containing tooth / dendrite development / negative regulation of cell cycle / mesoderm formation / positive regulation of SMAD protein signal transduction / epithelial to mesenchymal transition / negative regulation of Notch signaling pathway / positive regulation of osteoblast differentiation / positive regulation of bone mineralization / BMP signaling pathway / positive regulation of epithelial to mesenchymal transition / Transcriptional regulation of brown and beige adipocyte differentiation by EBF2 / neuron projection morphogenesis / positive regulation of brown fat cell differentiation / positive regulation of neuron differentiation / cytokine activity / axon guidance / growth factor activity / skeletal system development / negative regulation of neurogenesis / response to peptide hormone / osteoblast differentiation / response to estradiol / heparin binding / heart development / extracellular matrix / cellular response to hypoxia / vesicle / positive regulation of apoptotic process / negative regulation of DNA-templated transcription / positive regulation of gene expression / positive regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II / : / extracellular region Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.8 Å X-RAY DIFFRACTION / Resolution: 2.8 Å | ||||||
 Authors Authors | Griffith, D.L. / Scott, D.L. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 1996 Journal: Proc.Natl.Acad.Sci.USA / Year: 1996Title: Three-dimensional structure of recombinant human osteogenic protein 1: structural paradigm for the transforming growth factor beta superfamily. Authors: Griffith, D.L. / Keck, P.C. / Sampath, T.K. / Rueger, D.C. / Carlson, W.D. #1:  Journal: J.Mol.Biol. / Year: 1994 Journal: J.Mol.Biol. / Year: 1994Title: Crystallization and Preliminary Crystallographic Data of Recombinant Human Osteogenic Protein-1 (Hop-1) Authors: Griffith, D.L. / Oppermann, H. / Rueger, D.C. / Sampath, T.K. / Tucker, R.F. / Carlson, W.D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1bmp.cif.gz 1bmp.cif.gz | 33.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1bmp.ent.gz pdb1bmp.ent.gz | 22.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1bmp.json.gz 1bmp.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bm/1bmp https://data.pdbj.org/pub/pdb/validation_reports/bm/1bmp ftp://data.pdbj.org/pub/pdb/validation_reports/bm/1bmp ftp://data.pdbj.org/pub/pdb/validation_reports/bm/1bmp | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 15699.730 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) Homo sapiens (human)Description: REFERENCE, T.K. SAMPATH, ET AL. (1992) J. BIOL. CHEM. 267, 20452-20362 Gene: HOP-1 CDNA / Organ: OVARY / Gene (production host): HOP-1 CDNA / Production host:  |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.83 Å3/Da / Density % sol: 60 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal | *PLUS Density % sol: 62 % | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 22 ℃ / pH: 5 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Wavelength: 1.5418 |
|---|---|
| Detector | Type: RIGAKU / Detector: IMAGE PLATE / Date: Nov 30, 1993 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Num. obs: 5502 / % possible obs: 87 % / Observed criterion σ(I): 2 / Redundancy: 2.6 % / Rmerge(I) obs: 0.067 |
| Reflection | *PLUS Highest resolution: 2.8 Å |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.8→10 Å / σ(F): 2 Details: LOOP REGION (RESIDUES 118 - 122) IS DISORDERED AND MODELED STEREOCHEMICALLY. NOTE THAT RESIDUE 59 IS DESCRIBED AS TRANS IN THE PAPER CITED ON JRNL RECORDS ABOVE BUT THE CURRENT MODEL ...Details: LOOP REGION (RESIDUES 118 - 122) IS DISORDERED AND MODELED STEREOCHEMICALLY. NOTE THAT RESIDUE 59 IS DESCRIBED AS TRANS IN THE PAPER CITED ON JRNL RECORDS ABOVE BUT THE CURRENT MODEL PRESENTED IN THIS ENTRY HAS RESIDUE 59 AS CIS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 28.48 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.25 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.8→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: PROFFT / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.227 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj


