Entry Database : PDB / ID : 4leeTitle Structure of the Als3 adhesin from Candida albicans, residues 1-313 (mature sequence), triple mutant in the binding cavity: K59M, A116V, Y301F Agglutinin-like protein 3 Keywords / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Candida albicans (yeast)Method / / / Resolution : 3 Å Authors Lin, J. / Garnett, J.A. / Cota, E. Journal : J.Biol.Chem. / Year : 2014Title : The Peptide-binding Cavity Is Essential for Als3-mediated Adhesion of Candida albicans to Human Cells.Authors : Lin, J. / Oh, S.H. / Jones, R. / Garnett, J.A. / Salgado, P.S. / Rusnakova, S. / Matthews, S.J. / Hoyer, L.L. / Cota, E. History Deposition Jun 25, 2013 Deposition site / Processing site Revision 1.0 May 14, 2014 Provider / Type Revision 1.1 Jun 18, 2014 Group Revision 1.2 Jul 16, 2014 Group Revision 1.3 Sep 20, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / struct_ncs_dom_lim / struct_ref_seq_dif Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_ncs_dom_lim.beg_auth_comp_id / _struct_ncs_dom_lim.beg_label_asym_id / _struct_ncs_dom_lim.beg_label_comp_id / _struct_ncs_dom_lim.beg_label_seq_id / _struct_ncs_dom_lim.end_auth_comp_id / _struct_ncs_dom_lim.end_label_asym_id / _struct_ncs_dom_lim.end_label_comp_id / _struct_ncs_dom_lim.end_label_seq_id / _struct_ref_seq_dif.details Revision 1.4 Nov 27, 2024 Group / Category / pdbx_modification_feature
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Candida albicans (yeast)
Candida albicans (yeast) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3 Å
MOLECULAR REPLACEMENT / Resolution: 3 Å  Authors
Authors Citation
Citation Journal: J.Biol.Chem. / Year: 2014
Journal: J.Biol.Chem. / Year: 2014 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4lee.cif.gz
4lee.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4lee.ent.gz
pdb4lee.ent.gz PDB format
PDB format 4lee.json.gz
4lee.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/le/4lee
https://data.pdbj.org/pub/pdb/validation_reports/le/4lee ftp://data.pdbj.org/pub/pdb/validation_reports/le/4lee
ftp://data.pdbj.org/pub/pdb/validation_reports/le/4lee
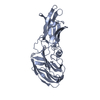
 Links
Links Assembly
Assembly




 Components
Components Candida albicans (yeast) / Gene: ALD8, ALS3 / Plasmid: PET32 XA/LIC / Production host:
Candida albicans (yeast) / Gene: ALD8, ALS3 / Plasmid: PET32 XA/LIC / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  Diamond
Diamond  / Beamline: I24 / Wavelength: 0.979 Å
/ Beamline: I24 / Wavelength: 0.979 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj