[English] 日本語
 Yorodumi
Yorodumi- PDB-1b6m: HIV-1 PROTEASE COMPLEXED WITH MACROCYCLIC PEPTIDOMIMETIC INHIBITOR 6 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1b6m | ||||||
|---|---|---|---|---|---|---|---|
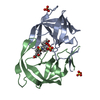
| Title | HIV-1 PROTEASE COMPLEXED WITH MACROCYCLIC PEPTIDOMIMETIC INHIBITOR 6 | ||||||
 Components Components | RETROPEPSIN | ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE INHIBITOR / COMPLEX (ACID PROTEINASE-PEPTIDE) / HYDROLASE-HYDROLASE INHIBITOR COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationHIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding ...HIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / viral penetration into host nucleus / host multivesicular body / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / host cell / viral nucleocapsid / DNA recombination / DNA-directed DNA polymerase / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / symbiont-mediated suppression of host gene expression / viral translational frameshifting / symbiont entry into host cell / lipid binding / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / proteolysis / DNA binding / zinc ion binding Similarity search - Function | ||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | ||||||
| Method |  X-RAY DIFFRACTION / OTHER / Resolution: 1.85 Å X-RAY DIFFRACTION / OTHER / Resolution: 1.85 Å | ||||||
 Authors Authors | Martin, J.L. / Begun, J. / Schindeler, A. / Wickramasinghe, W.A. / Alewood, D. / Alewood, P.F. / Bergman, D.A. / Brinkworth, R.I. / Abbenante, G. / March, D.R. ...Martin, J.L. / Begun, J. / Schindeler, A. / Wickramasinghe, W.A. / Alewood, D. / Alewood, P.F. / Bergman, D.A. / Brinkworth, R.I. / Abbenante, G. / March, D.R. / Reid, R.C. / Fairlie, D.P. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: Molecular recognition of macrocyclic peptidomimetic inhibitors by HIV-1 protease. Authors: Martin, J.L. / Begun, J. / Schindeler, A. / Wickramasinghe, W.A. / Alewood, D. / Alewood, P.F. / Bergman, D.A. / Brinkworth, R.I. / Abbenante, G. / March, D.R. / Reid, R.C. / Fairlie, D.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1b6m.cif.gz 1b6m.cif.gz | 56.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1b6m.ent.gz pdb1b6m.ent.gz | 39.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1b6m.json.gz 1b6m.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b6/1b6m https://data.pdbj.org/pub/pdb/validation_reports/b6/1b6m ftp://data.pdbj.org/pub/pdb/validation_reports/b6/1b6m ftp://data.pdbj.org/pub/pdb/validation_reports/b6/1b6m | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1b6jC  1b6kC  1b6lC  1b6pC  1z1hC  1z1rC  1cpiS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 10765.687 Da / Num. of mol.: 2 Mutation: GLN7LYS, LEU33ILE, CYS67ABA, CYS95ABA, GLN107LYS, LEU133ILE, CYS167ABA, CYS195ABA Source method: isolated from a natural source / Details: CYS RESIDUES REPLACED WITH ABA / Source: (natural)   Human immunodeficiency virus 1 / Genus: Lentivirus / References: UniProt: P03369, HIV-1 retropepsin Human immunodeficiency virus 1 / Genus: Lentivirus / References: UniProt: P03369, HIV-1 retropepsin#2: Chemical | #3: Chemical | ChemComp-PI6 / [ | 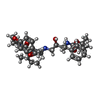  Type: peptide-like, Peptide-like / Class: Inhibitor / Mass: 596.757 Da / Num. of mol.: 1 / Source method: obtained synthetically / Formula: C33H48N4O6 Type: peptide-like, Peptide-like / Class: Inhibitor / Mass: 596.757 Da / Num. of mol.: 1 / Source method: obtained synthetically / Formula: C33H48N4O6References: tert-butyl [(1S,2S)-1-benzyl-2-hydroxy-3-{[(8S,11R)-8-[(1R)-1-methylpropyl]-7,10-dioxo-2-oxa-6,9-diazabicyclo[11.2.2]heptadeca-1(15),13,16-trien-11-yl]amino}propyl]carbamate #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.18 Å3/Da / Density % sol: 43.46 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 5.5 Details: 0.1 M ACETATE BUFFER PH 5.5 AND 30-60% AMMONIUM SULFATE | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging dropDetails: the enzyme/inhibitor solution was mixed in a 1:1 ratio with precipitant solution | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 289 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS IIC / Detector: IMAGE PLATE / Date: May 1, 1995 |
| Radiation | Monochromator: YALE MIRRORS / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.76→50 Å / Num. obs: 17002 / % possible obs: 88 % / Observed criterion σ(I): 1 / Redundancy: 3.1 % / Biso Wilson estimate: 11.8 Å2 / Rsym value: 0.046 / Net I/σ(I): 12.3 |
| Reflection shell | Resolution: 1.76→1.82 Å / Mean I/σ(I) obs: 2.7 / Rsym value: 0.276 / % possible all: 71 |
| Reflection | *PLUS % possible obs: 87.5 % / Num. measured all: 52836 / Rmerge(I) obs: 0.046 |
| Reflection shell | *PLUS % possible obs: 71.2 % / Rmerge(I) obs: 0.276 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: OTHER Starting model: PDB ENTRY 1CPI Resolution: 1.85→8 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 10000000 / Data cutoff low absF: 0.001 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 22.7 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.85→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.85→1.96 Å / Rfactor Rfree error: 0.019 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.851 / Classification: refinement X-PLOR / Version: 3.851 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller















 PDBj
PDBj




