+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 193d | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | SOLUTION STRUCTURE OF A QUINOMYCIN BISINTERCALATOR-DNA COMPLEX | ||||||||||||
 Components Components |
| ||||||||||||
 Keywords Keywords | DNA/ANTIBIOTIC / BISINTERCALATOR / DEPSIPEPTIDE / QUINOXALINE / THIOACETAL / ANTIBIOTIC / ANTITUMOR / DNA-ANTIBIOTIC COMPLEX | ||||||||||||
| Function / homology | uk63052 / 3-HYDROXYQUINALDIC ACID / DNA Function and homology information Function and homology information | ||||||||||||
| Biological species |  STREPTOMYCES (bacteria) STREPTOMYCES (bacteria) | ||||||||||||
| Method | SOLUTION NMR / MOLECULAR DYNAMICS, MATRIX RELAXATION | ||||||||||||
 Authors Authors | Chen, H. / Patel, D.J. | ||||||||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1995 Journal: J.Mol.Biol. / Year: 1995Title: Solution Structure of a Quinomycin Bisintercalator-DNA Complex. Authors: Chen, H. / Patel, D.J. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  193d.cif.gz 193d.cif.gz | 68.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb193d.ent.gz pdb193d.ent.gz | 52.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  193d.json.gz 193d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/93/193d https://data.pdbj.org/pub/pdb/validation_reports/93/193d ftp://data.pdbj.org/pub/pdb/validation_reports/93/193d ftp://data.pdbj.org/pub/pdb/validation_reports/93/193d | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Atom site foot note | 1: ALA D 2 - CYS D 3 MODEL 1 OMEGA = 0.00 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION 2: ALA C 2 - CYS C 3 MODEL 2 OMEGA = 0.00 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION 3: ALA D 2 - CYS D 3 MODEL 3 OMEGA = 0.00 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION 4: ALA D 2 - CYS D 3 MODEL 4 OMEGA = 359.98 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION 7: SER C 1 IS A D-SERINE. / 8: SER D 1 IS A D-SERINE. | |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 2426.617 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: CHEMICALLY SYNTHESIZED #2: Protein/peptide | | #3: Chemical | 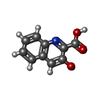  Type: Cyclic depsipeptide / Class: Antibiotic / Mass: 189.167 Da / Num. of mol.: 2 / Source method: obtained synthetically / Formula: C10H7NO3 Type: Cyclic depsipeptide / Class: Antibiotic / Mass: 189.167 Da / Num. of mol.: 2 / Source method: obtained synthetically / Formula: C10H7NO3Details: QUINOMYCIN UK63052 IS A BICYCLIC OCTADEPSIPEPTIDE. BICYCLIZATION IS ACHIEVED BY LINKING THE N- AND THE C- TERMINI, AND A THIOACETAL BOND BETWEEN RESIDUES 3 AND 7. THE TWO HYDROXYQUINOXALINE ...Details: QUINOMYCIN UK63052 IS A BICYCLIC OCTADEPSIPEPTIDE. BICYCLIZATION IS ACHIEVED BY LINKING THE N- AND THE C- TERMINI, AND A THIOACETAL BOND BETWEEN RESIDUES 3 AND 7. THE TWO HYDROXYQUINOXALINE CHROMOPHORES ARE LINKED TO THE D-SERINE RESIDUES, RESIDUES 1 AND 5. References: uk63052 Compound details | QUINOMYCIN UK63052 IS A BICYCLIC OCTADEPSIPEPTIDE, A MEMBER OF THE QUINOXALINE CLASS OF ANTIBIOTICS. ...QUINOMYCIN | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Details | Solvent system: D2O |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
- Processing
Processing
| Software |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| NMR software | Name:  X-PLOR / Developer: BRUNGER / Classification: refinement X-PLOR / Developer: BRUNGER / Classification: refinement | ||||||||
| Refinement | Method: MOLECULAR DYNAMICS, MATRIX RELAXATION / Software ordinal: 1 Details: TWO STARTING STRUCTURES WERE GENERATED BY MANUALLY DOCKING THE DRUG ONTO A- AND B-FORM DNA USING INSIGHT II AND WERE SUBSEQUENTLY REFINED BY DISTANCE- RESTRAINED MOLECULAR DYNAMICS USING A ...Details: TWO STARTING STRUCTURES WERE GENERATED BY MANUALLY DOCKING THE DRUG ONTO A- AND B-FORM DNA USING INSIGHT II AND WERE SUBSEQUENTLY REFINED BY DISTANCE- RESTRAINED MOLECULAR DYNAMICS USING A SET OF INTER-PROTON DISTANCE RESTRAINTS DERIVED FROM THE NMR DATA. TWO INITIAL VELOCITY SEEDS WERE USED FOR EACH STARTING STRUCTURE WHICH YIELDS FOUR DISTANCE-REFINED STRUCTURES. THEY WERE REFINED FURTHER USING RELAXATION-MATRIX BASED NOE INTENSITY-RESTRAINED MOLECULAR DYNAMICS. THE FINAL FOUR STRUCTURES WERE OBTAINED BY TAKING THE AVERAGE COORDINATES OF THE LAST 0.5 PS OF THE DYNAMICS DURING RELAXATION MATRIX REFINEMENT AND ENERGY MINIMIZED. THE R(1/6) VALUE WAS USED TO REFINE THE STRUCTURE DURING RELAXATION MATRIX REFINEMENT. THE SUMMATIONS RUN THROUGH ALL OBSERVED, QUANTIFIABLE NOE CROSSPEAKS IN NOESY SPECTRA RECORDED IN D2O AND MIXING TIMES OF 30, 60, 120 AND 180 MS. THE R(1/6) FACTOR AND THE RMS DEVIATIONS FROM IDEAL GEOMETRY FOR THE FOUR FINAL STRUCTURES ARE: MODEL1 MODEL2 MODEL3 MODEL4 R(1/6) FACTOR 0.023 0.024 0.024 0.026 BOND (ANG) 0.011 0.011 0.011 0.011 ANGLES (DEG) 3.616 3.657 3.700 3.725 IMPROPERS (DEG) 0.274 0.285 0.241 0.298 | ||||||||
| NMR ensemble | Conformer selection criteria: all calculated structures submitted Conformers calculated total number: 4 / Conformers submitted total number: 4 |
 Movie
Movie Controller
Controller













 PDBj
PDBj

