[English] 日本語
 Yorodumi
Yorodumi- EMDB-9908: Structure of PSI-isiA supercomplex from Thermosynechococcus vulcanus -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9908 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of PSI-isiA supercomplex from Thermosynechococcus vulcanus | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Electron Transport / PHOTOSYNTHESIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationphotosystem I reaction center / photosystem I / photosynthetic electron transport in photosystem I / photosystem I / plasma membrane-derived thylakoid membrane / chlorophyll binding / photosynthesis / 4 iron, 4 sulfur cluster binding / electron transfer activity / oxidoreductase activity ...photosystem I reaction center / photosystem I / photosynthetic electron transport in photosystem I / photosystem I / plasma membrane-derived thylakoid membrane / chlorophyll binding / photosynthesis / 4 iron, 4 sulfur cluster binding / electron transfer activity / oxidoreductase activity / magnesium ion binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Thermosynechococcus vulcanus (bacteria) Thermosynechococcus vulcanus (bacteria) | |||||||||
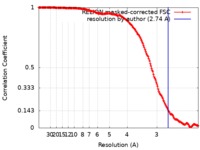
| Method | single particle reconstruction / cryo EM / Resolution: 2.74 Å | |||||||||
 Authors Authors | Akita F / Nagao R | |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2020 Journal: Commun Biol / Year: 2020Title: Structure of a cyanobacterial photosystem I surrounded by octadecameric IsiA antenna proteins. Authors: Fusamichi Akita / Ryo Nagao / Koji Kato / Yoshiki Nakajima / Makio Yokono / Yoshifumi Ueno / Takehiro Suzuki / Naoshi Dohmae / Jian-Ren Shen / Seiji Akimoto / Naoyuki Miyazaki /  Abstract: Iron-stress induced protein A (IsiA) is a chlorophyll-binding membrane-spanning protein in photosynthetic prokaryote cyanobacteria, and is associated with photosystem I (PSI) trimer cores, but its ...Iron-stress induced protein A (IsiA) is a chlorophyll-binding membrane-spanning protein in photosynthetic prokaryote cyanobacteria, and is associated with photosystem I (PSI) trimer cores, but its structural and functional significance in light harvesting remains unclear. Here we report a 2.7-Å resolution cryo-electron microscopic structure of a supercomplex between PSI core trimer and IsiA from a thermophilic cyanobacterium Thermosynechococcus vulcanus. The structure showed that 18 IsiA subunits form a closed ring surrounding a PSI trimer core. Detailed arrangement of pigments within the supercomplex, as well as molecular interactions between PSI and IsiA and among IsiAs, were resolved. Time-resolved fluorescence spectra of the PSI-IsiA supercomplex showed clear excitation-energy transfer from IsiA to PSI, strongly indicating that IsiA functions as an energy donor, but not an energy quencher, in the supercomplex. These structural and spectroscopic findings provide important insights into the excitation-energy-transfer and subunit assembly mechanisms in the PSI-IsiA supercomplex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9908.map.gz emd_9908.map.gz | 36.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9908-v30.xml emd-9908-v30.xml emd-9908.xml emd-9908.xml | 27.1 KB 27.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9908_fsc.xml emd_9908_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_9908.png emd_9908.png | 102.3 KB | ||
| Filedesc metadata |  emd-9908.cif.gz emd-9908.cif.gz | 8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9908 http://ftp.pdbj.org/pub/emdb/structures/EMD-9908 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9908 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9908 | HTTPS FTP |
-Related structure data
| Related structure data |  6k33MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9908.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9908.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.113 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
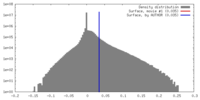
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : PSI-isiA
+Supramolecule #1: PSI-isiA
+Macromolecule #1: Photosystem I P700 chlorophyll a apoprotein A1
+Macromolecule #2: Photosystem I P700 chlorophyll a apoprotein A2
+Macromolecule #3: Photosystem I iron-sulfur center
+Macromolecule #4: Photosystem I reaction center subunit II
+Macromolecule #5: Photosystem I reaction center subunit IV
+Macromolecule #6: Photosystem I reaction center subunit III
+Macromolecule #7: Photosystem I reaction center subunit VIII
+Macromolecule #8: Photosystem I reaction center subunit IX
+Macromolecule #9: Photosystem I reaction center subunit PsaK
+Macromolecule #10: Photosystem I reaction center subunit XI
+Macromolecule #11: Photosystem I reaction center subunit XII
+Macromolecule #12: Photosystem I reaction center subunit psaX
+Macromolecule #13: Iron stress in-duced protein A
+Macromolecule #14: CHLOROPHYLL A ISOMER
+Macromolecule #15: CHLOROPHYLL A
+Macromolecule #16: PHYLLOQUINONE
+Macromolecule #17: IRON/SULFUR CLUSTER
+Macromolecule #18: BETA-CAROTENE
+Macromolecule #19: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
+Macromolecule #20: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #21: CALCIUM ION
+Macromolecule #22: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7 Component:
| |||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 30 sec. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)