+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0524 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of the photosystem I IsiA super-complex | |||||||||
 Map data Map data | PSI-IsiA super-complex from cyanobacteria | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Photosystem / Antenna / Chlorophyll / Membrane / PHOTOSYNTHESIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationcellular response to iron ion starvation / thylakoid / photosystem I reaction center / photosystem I / photosynthetic electron transport in photosystem I / photosynthetic electron transport chain / photosystem I / plasma membrane-derived thylakoid membrane / chlorophyll binding / photosynthesis ...cellular response to iron ion starvation / thylakoid / photosystem I reaction center / photosystem I / photosynthetic electron transport in photosystem I / photosynthetic electron transport chain / photosystem I / plasma membrane-derived thylakoid membrane / chlorophyll binding / photosynthesis / 4 iron, 4 sulfur cluster binding / electron transfer activity / oxidoreductase activity / magnesium ion binding / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.48 Å | |||||||||
 Authors Authors | Toporik H / Li J | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2019 Journal: Nat Struct Mol Biol / Year: 2019Title: The structure of the stress-induced photosystem I-IsiA antenna supercomplex. Authors: Hila Toporik / Jin Li / Dewight Williams / Po-Lin Chiu / Yuval Mazor /  Abstract: Photochemical conversion in oxygenic photosynthesis takes place in two large protein-pigment complexes named photosystem II and photosystem I (PSII and PSI, respectively). Photosystems associate with ...Photochemical conversion in oxygenic photosynthesis takes place in two large protein-pigment complexes named photosystem II and photosystem I (PSII and PSI, respectively). Photosystems associate with antennae in vivo to increase the size of photosynthetic units to hundreds or thousands of pigments. Regulation of the interactions between antennae and photosystems allows photosynthetic organisms to adapt to their environment. In low-iron environments, cyanobacteria express IsiA, a PSI antenna, critical to their survival. Here we describe the structure of the PSI-IsiA complex isolated from the mesophilic cyanobacterium Synechocystis sp. PCC 6803. This 2-MDa photosystem-antenna supercomplex structure reveals more than 700 pigments coordinated by 51 subunits, as well as the mechanisms facilitating the self-assembly and association of IsiA with multiple PSI assemblies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0524.map.gz emd_0524.map.gz | 39.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0524-v30.xml emd-0524-v30.xml emd-0524.xml emd-0524.xml | 25.1 KB 25.1 KB | Display Display |  EMDB header EMDB header |
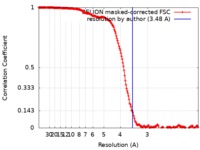
| FSC (resolution estimation) |  emd_0524_fsc.xml emd_0524_fsc.xml | 17.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_0524.png emd_0524.png | 78.6 KB | ||
| Filedesc metadata |  emd-0524.cif.gz emd-0524.cif.gz | 7.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0524 http://ftp.pdbj.org/pub/emdb/structures/EMD-0524 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0524 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0524 | HTTPS FTP |
-Related structure data
| Related structure data |  6nwaMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0524.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0524.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PSI-IsiA super-complex from cyanobacteria | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.024 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : PSI-IsiA
+Supramolecule #1: PSI-IsiA
+Macromolecule #1: Photosystem I P700 chlorophyll a apoprotein A1
+Macromolecule #2: Photosystem I P700 chlorophyll a apoprotein A2
+Macromolecule #3: Photosystem I iron-sulfur center
+Macromolecule #4: Photosystem I reaction center subunit II
+Macromolecule #5: Photosystem I reaction center subunit IV
+Macromolecule #6: Photosystem I reaction center subunit III
+Macromolecule #7: Photosystem I reaction center subunit VIII
+Macromolecule #8: Photosystem I reaction center subunit IX
+Macromolecule #9: Photosystem I reaction center subunit XI
+Macromolecule #10: Photosystem I reaction center subunit XII
+Macromolecule #11: Iron stress-induced chlorophyll-binding protein
+Macromolecule #12: Photosystem I reaction center subunit PsaK 1
+Macromolecule #13: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
+Macromolecule #14: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #15: CHLOROPHYLL A ISOMER
+Macromolecule #16: CHLOROPHYLL A
+Macromolecule #17: PHYLLOQUINONE
+Macromolecule #18: IRON/SULFUR CLUSTER
+Macromolecule #19: BETA-CAROTENE
+Macromolecule #20: DODECYL-BETA-D-MALTOSIDE
+Macromolecule #21: CALCIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 16.43 |
|---|---|
| Output model |  PDB-6nwa: |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)