+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9572 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the large subunit of the chloro-ribosome | |||||||||
 Map data Map data | Chloro-ribosome LSU (50S) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cryo-EM / ribosome / chloro-ribosome | |||||||||
| Function / homology |  Function and homology information Function and homology informationplastid translation / chloroplast envelope / mitochondrial large ribosomal subunit / mitochondrial translation / chloroplast thylakoid membrane / chloroplast / DNA-templated transcription termination / large ribosomal subunit / transferase activity / 5S rRNA binding ...plastid translation / chloroplast envelope / mitochondrial large ribosomal subunit / mitochondrial translation / chloroplast thylakoid membrane / chloroplast / DNA-templated transcription termination / large ribosomal subunit / transferase activity / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / mitochondrion / RNA binding Similarity search - Function | |||||||||
| Biological species |  Spinacia oleracea (spinach) Spinacia oleracea (spinach) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Ahmed T / Yin Z | |||||||||
| Funding support |  Singapore, 1 items Singapore, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2016 Journal: Sci Rep / Year: 2016Title: Cryo-EM structure of the large subunit of the spinach chloroplast ribosome. Authors: Tofayel Ahmed / Zhan Yin / Shashi Bhushan /  Abstract: Protein synthesis in the chloroplast is mediated by the chloroplast ribosome (chloro-ribosome). Overall architecture of the chloro-ribosome is considerably similar to the Escherichia coli (E. coli) ...Protein synthesis in the chloroplast is mediated by the chloroplast ribosome (chloro-ribosome). Overall architecture of the chloro-ribosome is considerably similar to the Escherichia coli (E. coli) ribosome but certain differences are evident. The chloro-ribosome proteins are generally larger because of the presence of chloroplast-specific extensions in their N- and C-termini. The chloro-ribosome harbours six plastid-specific ribosomal proteins (PSRPs); four in the small subunit and two in the large subunit. Deletions and insertions occur throughout the rRNA sequence of the chloro-ribosome (except for the conserved peptidyl transferase center region) but the overall length of the rRNAs do not change significantly, compared to the E. coli. Although, recent advancements in cryo-electron microscopy (cryo-EM) have provided detailed high-resolution structures of ribosomes from many different sources, a high-resolution structure of the chloro-ribosome is still lacking. Here, we present a cryo-EM structure of the large subunit of the chloro-ribosome from spinach (Spinacia oleracea) at an average resolution of 3.5 Å. High-resolution map enabled us to localize and model chloro-ribosome proteins, chloroplast-specific protein extensions, two PSRPs (PSRP5 and 6) and three rRNA molecules present in the chloro-ribosome. Although comparable to E. coli, the polypeptide tunnel and the tunnel exit site show chloroplast-specific features. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9572.map.gz emd_9572.map.gz | 11.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9572-v30.xml emd-9572-v30.xml emd-9572.xml emd-9572.xml | 49.8 KB 49.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9572.png emd_9572.png | 63.7 KB | ||
| Filedesc metadata |  emd-9572.cif.gz emd-9572.cif.gz | 11.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9572 http://ftp.pdbj.org/pub/emdb/structures/EMD-9572 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9572 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9572 | HTTPS FTP |
-Related structure data
| Related structure data |  5h1sMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9572.map.gz / Format: CCP4 / Size: 115.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9572.map.gz / Format: CCP4 / Size: 115.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Chloro-ribosome LSU (50S) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.28 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
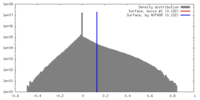
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Chloro-ribosome 50S
+Supramolecule #1: Chloro-ribosome 50S
+Macromolecule #1: 23S rRNA
+Macromolecule #2: Spinach chloroplast 4.5S rRNA
+Macromolecule #3: 5S rRNA
+Macromolecule #4: 50S ribosomal protein L13, chloroplastic
+Macromolecule #5: 50S ribosomal protein L14, chloroplastic
+Macromolecule #6: 50S ribosomal protein L15
+Macromolecule #7: 50S ribosomal protein L16, chloroplastic
+Macromolecule #8: 50S ribosomal protein L17
+Macromolecule #9: 50S ribosomal protein L18
+Macromolecule #10: 50S ribosomal protein L19, chloroplastic
+Macromolecule #11: 50S ribosomal protein L20, chloroplastic
+Macromolecule #12: 50S ribosomal protein L21, chloroplastic
+Macromolecule #13: 50S ribosomal protein L22, chloroplastic
+Macromolecule #14: 50S ribosomal protein L23, chloroplastic
+Macromolecule #15: 50S ribosomal protein L24, chloroplastic
+Macromolecule #16: 50S ribosomal protein L27
+Macromolecule #17: 50S ribosomal protein L28
+Macromolecule #18: 50S ribosomal protein L29
+Macromolecule #19: 50S ribosomal protein L2, chloroplastic
+Macromolecule #20: 50S ribosomal protein L32, chloroplastic
+Macromolecule #21: 50S ribosomal protein L33, chloroplastic
+Macromolecule #22: 50S ribosomal protein L34, chloroplastic
+Macromolecule #23: 50S ribosomal protein L35, chloroplastic
+Macromolecule #24: 50S ribosomal protein L36, chloroplastic
+Macromolecule #25: 50S ribosomal protein L3
+Macromolecule #26: 50S ribosomal protein L4, chloroplastic
+Macromolecule #27: 50S ribosomal protein L5, chloroplastic
+Macromolecule #28: 50S ribosomal protein L6
+Macromolecule #29: 50S ribosomal protein L9
+Macromolecule #30: 50S ribosomal protein 5 alpha, chloroplastic
+Macromolecule #31: 50S ribosomal protein L31
+Macromolecule #32: 50S ribosomal protein 6, chloroplastic
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Details: 20 mM Tris HCl, pH 7.6, 100 mM KCl, 10 mM MgOAc, 100 mM sucrose, 7 mM 2-mercaptoethanol, 1 unit/ml RNase inhibitor, 0.1% protease inhibitor |
|---|---|
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 1-7 / Number grids imaged: 1 / Number real images: 1590 / Average electron dose: 26.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 2.5 µm / Calibrated defocus min: 0.2 µm / Calibrated magnification: 109375 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.2 µm / Nominal magnification: 78000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)