+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9393 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron Cryo-Microscopy Of Chikungunya VLP | |||||||||
 Map data Map data | Chikungunya VLP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Chikungunya / virus-like particle / Structural Genomics / Center for Structural Genomics of Infectious Diseases / CSGID / VIRUS LIKE PARTICLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationtogavirin / T=4 icosahedral viral capsid / symbiont-mediated suppression of host toll-like receptor signaling pathway / host cell cytoplasm / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / symbiont entry into host cell / virion attachment to host cell / host cell nucleus / host cell plasma membrane ...togavirin / T=4 icosahedral viral capsid / symbiont-mediated suppression of host toll-like receptor signaling pathway / host cell cytoplasm / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / symbiont entry into host cell / virion attachment to host cell / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity / proteolysis / RNA binding / membrane Similarity search - Function | |||||||||
| Biological species |  Chikungunya virus (strain 37997) / Chikungunya virus (strain 37997) /   Chikungunya virus Chikungunya virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.16 Å | |||||||||
 Authors Authors | Basore K / Fremont DH | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2019 Journal: Cell / Year: 2019Title: Cryo-EM Structure of Chikungunya Virus in Complex with the Mxra8 Receptor. Authors: Katherine Basore / Arthur S Kim / Christopher A Nelson / Rong Zhang / Brittany K Smith / Carla Uranga / Lo Vang / Ming Cheng / Michael L Gross / Jonathan Smith / Michael S Diamond / Daved H Fremont /  Abstract: Mxra8 is a receptor for multiple arthritogenic alphaviruses that cause debilitating acute and chronic musculoskeletal disease in humans. Herein, we present a 2.2 Å resolution X-ray crystal ...Mxra8 is a receptor for multiple arthritogenic alphaviruses that cause debilitating acute and chronic musculoskeletal disease in humans. Herein, we present a 2.2 Å resolution X-ray crystal structure of Mxra8 and 4 to 5 Å resolution cryo-electron microscopy reconstructions of Mxra8 bound to chikungunya (CHIKV) virus-like particles and infectious virus. The Mxra8 ectodomain contains two strand-swapped Ig-like domains oriented in a unique disulfide-linked head-to-head arrangement. Mxra8 binds by wedging into a cleft created by two adjacent CHIKV E2-E1 heterodimers in one trimeric spike and engaging a neighboring spike. Two binding modes are observed with the fully mature VLP, with one Mxra8 binding with unique contacts. Only the high-affinity binding mode was observed in the complex with infectious CHIKV, as viral maturation and E3 occupancy appear to influence receptor binding-site usage. Our studies provide insight into how Mxra8 binds CHIKV and creates a path for developing alphavirus entry inhibitors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9393.map.gz emd_9393.map.gz | 590.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9393-v30.xml emd-9393-v30.xml emd-9393.xml emd-9393.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9393_fsc.xml emd_9393_fsc.xml | 19 KB | Display |  FSC data file FSC data file |
| Images |  emd_9393.png emd_9393.png | 247.6 KB | ||
| Filedesc metadata |  emd-9393.cif.gz emd-9393.cif.gz | 7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9393 http://ftp.pdbj.org/pub/emdb/structures/EMD-9393 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9393 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9393 | HTTPS FTP |
-Validation report
| Summary document |  emd_9393_validation.pdf.gz emd_9393_validation.pdf.gz | 784 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9393_full_validation.pdf.gz emd_9393_full_validation.pdf.gz | 783.5 KB | Display | |
| Data in XML |  emd_9393_validation.xml.gz emd_9393_validation.xml.gz | 17.3 KB | Display | |
| Data in CIF |  emd_9393_validation.cif.gz emd_9393_validation.cif.gz | 24 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9393 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9393 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9393 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9393 | HTTPS FTP |
-Related structure data
| Related structure data |  6nk5MC  9394C  9395C  6nk3C  6nk6C  6nk7C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9393.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9393.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Chikungunya VLP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.403 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
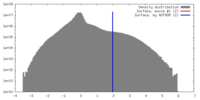
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Chikungunya virus
| Entire | Name:   Chikungunya virus Chikungunya virus |
|---|---|
| Components |
|
-Supramolecule #1: Chikungunya virus
| Supramolecule | Name: Chikungunya virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 / Details: Produced by PaxVax Corporation, Redwood CA, USA. / NCBI-ID: 37124 / Sci species name: Chikungunya virus / Sci species strain: 37997 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: Yes |
|---|
-Macromolecule #1: E1 glycoprotein
| Macromolecule | Name: E1 glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chikungunya virus (strain 37997) / Strain: 37997 Chikungunya virus (strain 37997) / Strain: 37997 |
| Molecular weight | Theoretical: 47.346715 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: YEHVTVIPNT VGVPYKTLVN RPGYSPMVLE MELQSVTLEP TLSLDYITCE YKTVIPSPYV KCCGTAECKD KSLPDYSCKV FTGVYPFMW GGAYCFCDAE NTQLSEAHVE KSESCKTEFA SAYRAHTASA SAKLRVLYQG NNITVAAYAN GDHAVTVKDA K FVVGPMSS ...String: YEHVTVIPNT VGVPYKTLVN RPGYSPMVLE MELQSVTLEP TLSLDYITCE YKTVIPSPYV KCCGTAECKD KSLPDYSCKV FTGVYPFMW GGAYCFCDAE NTQLSEAHVE KSESCKTEFA SAYRAHTASA SAKLRVLYQG NNITVAAYAN GDHAVTVKDA K FVVGPMSS AWTPFDNKIV VYKGDVYNMD YPPFGAGRPG QFGDIQSRTP ESKDVYANTQ LVLQRPAAGT VHVPYSQAPS GF KYWLKER GASLQHTAPF GCQIATNPVR AVNCAVGNIP ISIDIPDAAF TRVVDAPSVT DMSCEVPACT HSSDFGGVAI IKY TASKKG KCAVHSMTNA VTIREADVEV EGNSQLQISF STALASAEFR VQVCSTQVHC AAACHPPKDH IVNYPASHTT LGVQ DISTT AMSWVQKITG GVGLIVAVAA LILIVVLCVS FSRH UniProtKB: Structural polyprotein |
-Macromolecule #2: E2 glycoprotein
| Macromolecule | Name: E2 glycoprotein / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chikungunya virus (strain 37997) / Strain: 37997 Chikungunya virus (strain 37997) / Strain: 37997 |
| Molecular weight | Theoretical: 46.858312 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NFNVYKATRP YLAHCPDCGE GHSCHSPIAL ERIRNEATDG TLKIQVSLQI GIKTDDSHDW TKLRYMDSHT PADAERAGLL VRTSAPCTI TGTMGHFILA RCPKGETLTV GFTDSRKISH TCTHPFHHEP PVIGRERFHS RPQHGKELPC STYVQSTAAT A EEIEVHMP ...String: NFNVYKATRP YLAHCPDCGE GHSCHSPIAL ERIRNEATDG TLKIQVSLQI GIKTDDSHDW TKLRYMDSHT PADAERAGLL VRTSAPCTI TGTMGHFILA RCPKGETLTV GFTDSRKISH TCTHPFHHEP PVIGRERFHS RPQHGKELPC STYVQSTAAT A EEIEVHMP PDTPDRTLMT QQSGNVKITV NGQTVRYKCN CGGSNEGLTT TDKVINNCKI DQCHAAVTNH KNWQYNSPLV PR NAELGDR KGKIHIPFPL ANVTCRVPKA RNPTVTYGKN QVTMLLYPDH PTLLSYRNMG QEPNYHEEWV THKKEVTLTV PTE GLEVTW GNNEPYKYWP QMSTNGTAHG HPHEIILYYY ELYPTMTVVI VSVASFVLLS MVGTAVGMCV CARRRCITPY ELTP GATVP FLLSLLCCVR TTKA UniProtKB: Structural polyprotein |
-Macromolecule #3: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chikungunya virus (strain 37997) / Strain: 37997 Chikungunya virus (strain 37997) / Strain: 37997 |
| Molecular weight | Theoretical: 16.458701 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NDCIFEVKHE GKVMGYACLV GDKVMKPAHV KGTIDNADLA KLAFKRSSKY DLECAQIPVH MKSDASKFTH EKPEGYYNWH HGAVQYSGG RFTIPTGAGK PGDSGRPIFD NKGRVVAIVL GGANEGARTA LSVVTWNKDI VTKITPEGAE EW UniProtKB: Structural polyprotein |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 8 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.33 mg/mL | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.2 Component:
| ||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Support film - Film thickness: 12 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: OTHER / Details: 0.458 mbar.l/s O2 and 0.11 mbar.l/s H2 | ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average exposure time: 0.3 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT | ||||||||||||
| Output model |  PDB-6nk5: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)