+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8287 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM reconstruction of Neisseria meningitidis Type IV pilus | |||||||||
 Map data Map data | 3D cryo-EM reconstruction of Nm Type IV pilus | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | Fimbrial protein pilin / Pilin (bacterial filament) / Prokaryotic N-terminal methylation motif / Prokaryotic N-terminal methylation site / Pilin-like / pilus / cell adhesion / Pilin Function and homology information Function and homology information | |||||||||
| Biological species |  Neisseria meningitidis (bacteria) Neisseria meningitidis (bacteria) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 6.0 Å | |||||||||
 Authors Authors | Kolappan S / Coureuil M / Yu X / Nassif X / Craig L / Egelman EH | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2016 Journal: Nat Commun / Year: 2016Title: Structure of the Neisseria meningitidis Type IV pilus. Authors: Subramania Kolappan / Mathieu Coureuil / Xiong Yu / Xavier Nassif / Edward H Egelman / Lisa Craig /    Abstract: Neisseria meningitidis use Type IV pili (T4P) to adhere to endothelial cells and breach the blood brain barrier, causing cause fatal meningitis. T4P are multifunctional polymers of the major pilin ...Neisseria meningitidis use Type IV pili (T4P) to adhere to endothelial cells and breach the blood brain barrier, causing cause fatal meningitis. T4P are multifunctional polymers of the major pilin protein, which share a conserved hydrophobic N terminus that is a curved extended α-helix, α1, in X-ray crystal structures. Here we report a 1.44 Å crystal structure of the N. meningitidis major pilin PilE and a ∼6 Å cryo-electron microscopy reconstruction of the intact pilus, from which we built an atomic model for the filament. This structure reveals the molecular arrangement of the N-terminal α-helices in the filament core, including a melted central portion of α1 and a bridge of electron density consistent with a predicted salt bridge necessary for pilus assembly. This structure has important implications for understanding pilus biology. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8287.map.gz emd_8287.map.gz | 4.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8287-v30.xml emd-8287-v30.xml emd-8287.xml emd-8287.xml | 10.3 KB 10.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8287.png emd_8287.png | 138.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8287 http://ftp.pdbj.org/pub/emdb/structures/EMD-8287 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8287 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8287 | HTTPS FTP |
-Related structure data
| Related structure data |  5kuaMC  5jw8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8287.map.gz / Format: CCP4 / Size: 7.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8287.map.gz / Format: CCP4 / Size: 7.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D cryo-EM reconstruction of Nm Type IV pilus | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
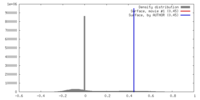
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : pilus
| Entire | Name: pilus |
|---|---|
| Components |
|
-Supramolecule #1: pilus
| Supramolecule | Name: pilus / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Neisseria meningitidis (bacteria) Neisseria meningitidis (bacteria) |
-Macromolecule #1: pilin
| Macromolecule | Name: pilin / type: other / ID: 1 / Classification: other |
|---|---|
| Source (natural) | Organism:  Neisseria meningitidis (bacteria) Neisseria meningitidis (bacteria) |
| Sequence | String: FTLIELMIVI AIVGILAAVA LPAYQDYTAR AQVSEAILLA EGQKSAVTEY YLNHGEWPGD NSSAGVATSA DIKGKYVQSV TVANGVITAQ MASSNVNNEI KSKKLSLWAK RQNGSVKWFC GQPVTRTTAT ATDVAAANGK TDDKINTKHL PSTCRDDSSA S |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 10.5 |
|---|---|
| Grid | Model: lacey / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: LACEY / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 297 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Number grids imaged: 1 / Number real images: 541 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 10.3 Å Applied symmetry - Helical parameters - Δ&Phi: 100.8 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 6.0 Å / Resolution method: OTHER / Number images used: 15586 |
|---|---|
| CTF correction | Software - Name: CTFFIND3 |
| Segment selection | Number selected: 15586 |
| Final angle assignment | Type: NOT APPLICABLE |
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-5kua: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)