[English] 日本語
 Yorodumi
Yorodumi- EMDB-8194: Cryo-EM structure of glutamate dehydrogenase at 1.8 A resolution -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8194 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of glutamate dehydrogenase at 1.8 A resolution | |||||||||
 Map data Map data | Glutamate dehydrogenase at 1.8 A resolution | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | glutamate dehydrogenase / small metabolic complex / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationL-glutamate dehydrogenase [NAD(P)+] activity / tricarboxylic acid metabolic process / glutamate dehydrogenase [NAD(P)+] / L-glutamate dehydrogenase (NADP+) activity / L-glutamate dehydrogenase (NAD+) activity / L-glutamate catabolic process / L-glutamine metabolic process / mitochondrial inner membrane / GTP binding / endoplasmic reticulum ...L-glutamate dehydrogenase [NAD(P)+] activity / tricarboxylic acid metabolic process / glutamate dehydrogenase [NAD(P)+] / L-glutamate dehydrogenase (NADP+) activity / L-glutamate dehydrogenase (NAD+) activity / L-glutamate catabolic process / L-glutamine metabolic process / mitochondrial inner membrane / GTP binding / endoplasmic reticulum / mitochondrion / ATP binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.8 Å | |||||||||
 Authors Authors | Merk A / Bartesaghi A | |||||||||
 Citation Citation |  Journal: Cell / Year: 2016 Journal: Cell / Year: 2016Title: Breaking Cryo-EM Resolution Barriers to Facilitate Drug Discovery. Authors: Alan Merk / Alberto Bartesaghi / Soojay Banerjee / Veronica Falconieri / Prashant Rao / Mindy I Davis / Rajan Pragani / Matthew B Boxer / Lesley A Earl / Jacqueline L S Milne / Sriram Subramaniam /  Abstract: Recent advances in single-particle cryoelecton microscopy (cryo-EM) are enabling generation of numerous near-atomic resolution structures for well-ordered protein complexes with sizes ≥ ∼200 kDa. ...Recent advances in single-particle cryoelecton microscopy (cryo-EM) are enabling generation of numerous near-atomic resolution structures for well-ordered protein complexes with sizes ≥ ∼200 kDa. Whether cryo-EM methods are equally useful for high-resolution structural analysis of smaller, dynamic protein complexes such as those involved in cellular metabolism remains an important question. Here, we present 3.8 Å resolution cryo-EM structures of the cancer target isocitrate dehydrogenase (93 kDa) and identify the nature of conformational changes induced by binding of the allosteric small-molecule inhibitor ML309. We also report 2.8-Å- and 1.8-Å-resolution structures of lactate dehydrogenase (145 kDa) and glutamate dehydrogenase (334 kDa), respectively. With these results, two perceived barriers in single-particle cryo-EM are overcome: (1) crossing 2 Å resolution and (2) obtaining structures of proteins with sizes < 100 kDa, demonstrating that cryo-EM can be used to investigate a broad spectrum of drug-target interactions and dynamic conformational states. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8194.map.gz emd_8194.map.gz | 45.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8194-v30.xml emd-8194-v30.xml emd-8194.xml emd-8194.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8194.png emd_8194.png | 134.7 KB | ||
| Filedesc metadata |  emd-8194.cif.gz emd-8194.cif.gz | 6 KB | ||
| Others |  emd_8194_additional.map.gz emd_8194_additional.map.gz | 45.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8194 http://ftp.pdbj.org/pub/emdb/structures/EMD-8194 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8194 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8194 | HTTPS FTP |
-Related structure data
| Related structure data |  5k12MC  8191C  8192C  8193C  5k0zC  5k10C  5k11C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8194.map.gz / Format: CCP4 / Size: 48.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8194.map.gz / Format: CCP4 / Size: 48.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Glutamate dehydrogenase at 1.8 A resolution | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.637 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
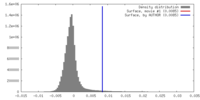
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Map sharpened using a B-factor of -90
| File | emd_8194_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map sharpened using a B-factor of -90 | ||||||||||||
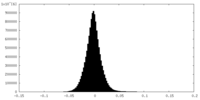
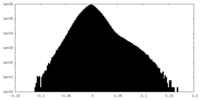
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Glutamate dehydrogenase
| Entire | Name: Glutamate dehydrogenase |
|---|---|
| Components |
|
-Supramolecule #1: Glutamate dehydrogenase
| Supramolecule | Name: Glutamate dehydrogenase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 334 KDa |
-Macromolecule #1: Glutamate dehydrogenase 1, mitochondrial
| Macromolecule | Name: Glutamate dehydrogenase 1, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO / EC number: glutamate dehydrogenase [NAD(P)+] |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 61.60891 KDa |
| Sequence | String: MYRYLGEALL LSRAGPAALG SASADSAALL GWARGQPAAA PQPGLVPPAR RHYSEAAADR EDDPNFFKMV EGFFDRGASI VEDKLVEDL KTRETEEQKR NRVRSILRII KPCNHVLSLS FPIRRDDGSW EVIEGYRAQH SQHRTPCKGG IRYSTDVSVD E VKALASLM ...String: MYRYLGEALL LSRAGPAALG SASADSAALL GWARGQPAAA PQPGLVPPAR RHYSEAAADR EDDPNFFKMV EGFFDRGASI VEDKLVEDL KTRETEEQKR NRVRSILRII KPCNHVLSLS FPIRRDDGSW EVIEGYRAQH SQHRTPCKGG IRYSTDVSVD E VKALASLM TYKCAVVDVP FGGAKAGVKI NPKNYTDNEL EKITRRFTME LAKKGFIGPG VDVPAPDMST GEREMSWIAD TY ASTIGHY DINAHACVTG KPISQGGIHG RISATGRGVF HGIENFINEA SYMSILGMTP GFGDKTFVVQ GFGNVGLHSM RYL HRFGAK CITVGESDGS IWNPDGIDPK ELEDFKLQHG TILGFPKAKI YEGSILEVDC DILIPAASEK QLTKSNAPRV KAKI IAEGA NGPTTPEADK IFLERNIMVI PDLYLNAGGV TVSYFEWLKN LNHVSYGRLT FKYERDSNYH LLMSVQESLE RKFGK HGGT IPIVPTAEFQ DRISGASEKD IVHSGLAYTM ERSARQIMRT AMKYNLGLDL RTAAYVNAIE KVFRVYNEAG VTFT UniProtKB: Glutamate dehydrogenase 1, mitochondrial |
-Macromolecule #2: water
| Macromolecule | Name: water / type: ligand / ID: 2 / Number of copies: 1086 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 6.8 / Component - Concentration: 100.0 nM / Component - Name: Potassium phosphate |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV Details: Plunged into liquiq ethane (FEI VITROBOT MARK IV). |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 79.6 K / Max: 79.8 K |
| Specialist optics | Energy filter - Name: GIF Quantum / Energy filter - Lower energy threshold: 0 eV / Energy filter - Upper energy threshold: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 3-9 / Number real images: 232 / Average exposure time: 0.4 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 78000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 215000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)