[English] 日本語
 Yorodumi
Yorodumi- EMDB-7096: Cryo-EM structure of the TMEM16A calcium-activated chloride chann... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7096 | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the TMEM16A calcium-activated chloride channel in LMNG | ||||||||||||||||||||||||||||||
 Map data Map data | sharpened map | ||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||
 Keywords Keywords | Chloride channel / TMEM16 family / MEMBRANE PROTEIN | ||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationglial cell projection elongation / trachea development / iodide transmembrane transporter activity / mucus secretion / iodide transport / intracellularly calcium-gated chloride channel activity / cellular response to peptide / Stimuli-sensing channels / voltage-gated chloride channel activity / calcium-activated cation channel activity ...glial cell projection elongation / trachea development / iodide transmembrane transporter activity / mucus secretion / iodide transport / intracellularly calcium-gated chloride channel activity / cellular response to peptide / Stimuli-sensing channels / voltage-gated chloride channel activity / calcium-activated cation channel activity / protein localization to membrane / chloride transport / detection of temperature stimulus involved in sensory perception of pain / chloride channel activity / chloride channel complex / chloride transmembrane transport / regulation of membrane potential / cellular response to heat / presynaptic membrane / phospholipase C-activating G protein-coupled receptor signaling pathway / apical plasma membrane / signaling receptor binding / external side of plasma membrane / glutamatergic synapse / protein homodimerization activity / nucleoplasm / metal ion binding / identical protein binding / plasma membrane Similarity search - Function | ||||||||||||||||||||||||||||||
| Biological species |  | ||||||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | ||||||||||||||||||||||||||||||
 Authors Authors | Dang S / Feng S | ||||||||||||||||||||||||||||||
| Funding support |  United States, 9 items United States, 9 items
| ||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2017 Journal: Nature / Year: 2017Title: Cryo-EM structures of the TMEM16A calcium-activated chloride channel. Authors: Shangyu Dang / Shengjie Feng / Jason Tien / Christian J Peters / David Bulkley / Marco Lolicato / Jianhua Zhao / Kathrin Zuberbühler / Wenlei Ye / Lijun Qi / Tingxu Chen / Charles S Craik / ...Authors: Shangyu Dang / Shengjie Feng / Jason Tien / Christian J Peters / David Bulkley / Marco Lolicato / Jianhua Zhao / Kathrin Zuberbühler / Wenlei Ye / Lijun Qi / Tingxu Chen / Charles S Craik / Yuh Nung Jan / Daniel L Minor / Yifan Cheng / Lily Yeh Jan /  Abstract: Calcium-activated chloride channels (CaCCs) encoded by TMEM16A control neuronal signalling, smooth muscle contraction, airway and exocrine gland secretion, and rhythmic movements of the ...Calcium-activated chloride channels (CaCCs) encoded by TMEM16A control neuronal signalling, smooth muscle contraction, airway and exocrine gland secretion, and rhythmic movements of the gastrointestinal system. To understand how CaCCs mediate and control anion permeation to fulfil these physiological functions, knowledge of the mammalian TMEM16A structure and identification of its pore-lining residues are essential. TMEM16A forms a dimer with two pores. Previous CaCC structural analyses have relied on homology modelling of a homologue (nhTMEM16) from the fungus Nectria haematococca that functions primarily as a lipid scramblase, as well as subnanometre-resolution electron cryo-microscopy. Here we present de novo atomic structures of the transmembrane domains of mouse TMEM16A in nanodiscs and in lauryl maltose neopentyl glycol as determined by single-particle electron cryo-microscopy. These structures reveal the ion permeation pore and represent different functional states. The structure in lauryl maltose neopentyl glycol has one Ca ion resolved within each monomer with a constricted pore; this is likely to correspond to a closed state, because a CaCC with a single Ca occupancy requires membrane depolarization in order to open (C.J.P. et al., manuscript submitted). The structure in nanodiscs has two Ca ions per monomer and its pore is in a closed conformation; this probably reflects channel rundown, which is the gradual loss of channel activity that follows prolonged CaCC activation in 1 mM Ca. Our mutagenesis and electrophysiological studies, prompted by analyses of the structures, identified ten residues distributed along the pore that interact with permeant anions and affect anion selectivity, as well as seven pore-lining residues that cluster near pore constrictions and regulate channel gating. Together, these results clarify the basis of CaCC anion conduction. | ||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7096.map.gz emd_7096.map.gz | 60 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7096-v30.xml emd-7096-v30.xml emd-7096.xml emd-7096.xml | 31.6 KB 31.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7096.png emd_7096.png | 54.1 KB | ||
| Filedesc metadata |  emd-7096.cif.gz emd-7096.cif.gz | 8 KB | ||
| Others |  emd_7096_additional.map.gz emd_7096_additional.map.gz emd_7096_half_map_1.map.gz emd_7096_half_map_1.map.gz emd_7096_half_map_2.map.gz emd_7096_half_map_2.map.gz | 48.6 MB 48.6 MB 48.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7096 http://ftp.pdbj.org/pub/emdb/structures/EMD-7096 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7096 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7096 | HTTPS FTP |
-Related structure data
| Related structure data |  6bgjMC  7095C  6bgiC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10124 (Title: Cryo-EM structure of the TMEM16A in LMNG / Data size: 244.2 EMPIAR-10124 (Title: Cryo-EM structure of the TMEM16A in LMNG / Data size: 244.2 Data #1: Particle stacks extracted from motion corrected micrographs of TMEM16A in LMNG [picked particles - single frame - processed] Data #2: Particle stacks extracted from motion corrected micrographs of TMEM16A bound with Fab in LMNG [picked particles - single frame - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7096.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7096.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.02 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
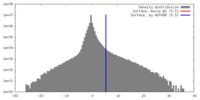
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unsharpened map
| File | emd_7096_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
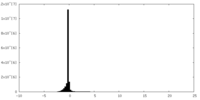
| Density Histograms |
-Half map: half map 1
| File | emd_7096_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
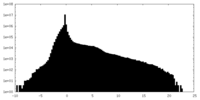
| Density Histograms |
-Half map: half map 2
| File | emd_7096_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
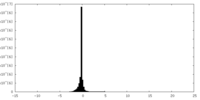
| Density Histograms |
- Sample components
Sample components
-Entire : TMEM16A Calcium-activated chloride channel
| Entire | Name: TMEM16A Calcium-activated chloride channel |
|---|---|
| Components |
|
-Supramolecule #1: TMEM16A Calcium-activated chloride channel
| Supramolecule | Name: TMEM16A Calcium-activated chloride channel / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Anoctamin-1
| Macromolecule | Name: Anoctamin-1 / type: protein_or_peptide / ID: 1 / Details: Calcium ions / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 105.603984 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRVPEKYSTL PAEDRSVHIV NICAIEDLGY LPSEGTLLNS LSVDPDAECK YGLYFRDGKR KVDYILVYHH KRASGSRTLA RRGLQNDMV LGTRSVRQDQ PLPGKGSPVD AGSPEVPMDY HEDDKRFRRE EYEGNLLEAG LELENDEDTK IHGVGFVKIH A PWHVLCRE ...String: MRVPEKYSTL PAEDRSVHIV NICAIEDLGY LPSEGTLLNS LSVDPDAECK YGLYFRDGKR KVDYILVYHH KRASGSRTLA RRGLQNDMV LGTRSVRQDQ PLPGKGSPVD AGSPEVPMDY HEDDKRFRRE EYEGNLLEAG LELENDEDTK IHGVGFVKIH A PWHVLCRE AEFLKLKMPT KKVYHISETR GLLKTINSVL QKITDPIQPK VAEHRPQTTK RLSYPFSREK QHLFDLTDRD SF FDSKTRS TIVYEILKRT TCTKAKYSMG ITSLLANGVY SAAYPLHDGD YEGDNVEFND RKLLYEEWAS YGVFYKYQPI DLV RKYFGE KVGLYFAWLG AYTQMLIPAS IVGVIVFLYG CATVDENIPS MEMCDQRYNI TMCPLCDKTC SYWKMSSACA TARA SHLFD NPATVFFSVF MALWAATFME HWKRKQMRLN YRWDLTGFEE EEDHPRAEYE ARVLEKSLRK ESRNKETDKV KLTWR DRFP AYFTNLVSII FMIAVTFAIV LGVIIYRIST AAALAMNSSP SVRSNIRVTV TATAVIINLV VIILLDEVYG CIARWL TKI EVPKTEKSFE ERLTFKAFLL KFVNSYTPIF YVAFFKGRFV GRPGDYVYIF RSFRMEECAP GGCLMELCIQ LSIIMLG KQ LIQNNLFEIG IPKMKKFIRY LKLRRQSPSD REEYVKRKQR YEVDFNLEPF AGLTPEYMEM IIQFGFVTLF VASFPLAP L FALLNNIIEI RLDAKKFVTE LRRPVAIRAK DIGIWYNILR GVGKLAVIIN AFVISFTSDF IPRLVYLYMY SQNGTMHGF VNHTLSSFNV SDFQNGTAPN DPLDLGYEVQ ICRYKDYREP PWSEHKYDIS KDFWAVLAAR LAFVIVFQNL VMFMSDFVDW VIPDIPKDI SQQIHKEKVL MVELSNSLEV LFQ UniProtKB: Anoctamin-1 |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 9 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 10 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.027 kPa | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III / Details: blot 6-8 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-40 / Number grids imaged: 2 / Number real images: 7882 / Average exposure time: 8.0 sec. / Average electron dose: 80.0 e/Å2 Details: 5326 images for TMEM16A alone in LMNG, 2556 images for TMEM16A with Fab bound in LMNG |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 1.9000000000000001 µm / Calibrated defocus min: 0.8 µm / Calibrated magnification: 29000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6bgj: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)