[English] 日本語
 Yorodumi
Yorodumi- EMDB-47959: Cryo-EM model of E. coli aspartate transcarbamoylase in the T-sta... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM model of E. coli aspartate transcarbamoylase in the T-state complexed with CP, CTP, and Mg2+ | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Complex / Allostery / ATCase / R-State / CTP / UTP / CP / Succinate / CYTOSOLIC PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationaspartate carbamoyltransferase complex / pyrimidine nucleotide biosynthetic process / aspartate carbamoyltransferase / aspartate carbamoyltransferase activity / amino acid metabolic process / amino acid binding / 'de novo' UMP biosynthetic process / 'de novo' pyrimidine nucleobase biosynthetic process / transferase activity / metal ion binding / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.85 Å | |||||||||||||||
 Authors Authors | Miller RC / Ando N | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation | Journal: Acta Crystallogr D Struct Biol / Year: 2019 Title: Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Authors: Dorothee Liebschner / Pavel V Afonine / Matthew L Baker / Gábor Bunkóczi / Vincent B Chen / Tristan I Croll / Bradley Hintze / Li Wei Hung / Swati Jain / Airlie J McCoy / Nigel W Moriarty ...Authors: Dorothee Liebschner / Pavel V Afonine / Matthew L Baker / Gábor Bunkóczi / Vincent B Chen / Tristan I Croll / Bradley Hintze / Li Wei Hung / Swati Jain / Airlie J McCoy / Nigel W Moriarty / Robert D Oeffner / Billy K Poon / Michael G Prisant / Randy J Read / Jane S Richardson / David C Richardson / Massimo D Sammito / Oleg V Sobolev / Duncan H Stockwell / Thomas C Terwilliger / Alexandre G Urzhumtsev / Lizbeth L Videau / Christopher J Williams / Paul D Adams /    Abstract: Diffraction (X-ray, neutron and electron) and electron cryo-microscopy are powerful methods to determine three-dimensional macromolecular structures, which are required to understand biological ...Diffraction (X-ray, neutron and electron) and electron cryo-microscopy are powerful methods to determine three-dimensional macromolecular structures, which are required to understand biological processes and to develop new therapeutics against diseases. The overall structure-solution workflow is similar for these techniques, but nuances exist because the properties of the reduced experimental data are different. Software tools for structure determination should therefore be tailored for each method. Phenix is a comprehensive software package for macromolecular structure determination that handles data from any of these techniques. Tasks performed with Phenix include data-quality assessment, map improvement, model building, the validation/rebuilding/refinement cycle and deposition. Each tool caters to the type of experimental data. The design of Phenix emphasizes the automation of procedures, where possible, to minimize repetitive and time-consuming manual tasks, while default parameters are chosen to encourage best practice. A graphical user interface provides access to many command-line features of Phenix and streamlines the transition between programs, project tracking and re-running of previous tasks. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_47959.map.gz emd_47959.map.gz | 141.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-47959-v30.xml emd-47959-v30.xml emd-47959.xml emd-47959.xml | 35.8 KB 35.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_47959_fsc.xml emd_47959_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_47959.png emd_47959.png | 84.2 KB | ||
| Filedesc metadata |  emd-47959.cif.gz emd-47959.cif.gz | 10 KB | ||
| Others |  emd_47959_additional_1.map.gz emd_47959_additional_1.map.gz emd_47959_half_map_1.map.gz emd_47959_half_map_1.map.gz emd_47959_half_map_2.map.gz emd_47959_half_map_2.map.gz | 11.3 MB 141 MB 140.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-47959 http://ftp.pdbj.org/pub/emdb/structures/EMD-47959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-47959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-47959 | HTTPS FTP |
-Related structure data
| Related structure data |  9eenMC  9eehC  9eejC  9eekC  9eelC  9eemC  9eeoC  9eepC  9eeqC  9eerC  9eesC  9eeuC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_47959.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_47959.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.069 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_47959_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_47959_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_47959_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : E. coli aspartate transcarbamoylase in the T-state complexed with...
| Entire | Name: E. coli aspartate transcarbamoylase in the T-state complexed with CP, CTP, and Mg2+ |
|---|---|
| Components |
|
-Supramolecule #1: E. coli aspartate transcarbamoylase in the T-state complexed with...
| Supramolecule | Name: E. coli aspartate transcarbamoylase in the T-state complexed with CP, CTP, and Mg2+ type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2, #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Aspartate carbamoyltransferase regulatory chain
| Macromolecule | Name: Aspartate carbamoyltransferase regulatory chain / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.945273 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AIKRGTVIDH IPAQIGFKLL SLFKLTETDQ RITIGLNLPS GEMGRKDLIK IENTFLSEDQ VDQLALYAPQ ATVNRIDNYE VVGKSRPSL PERIDNVLVC PNSNCISHAE PVSSSFAVRK RANDIALKCK YCEKEFSHNV VLAN UniProtKB: Aspartate carbamoyltransferase regulatory chain |
-Macromolecule #2: Aspartate carbamoyltransferase
| Macromolecule | Name: Aspartate carbamoyltransferase / type: protein_or_peptide / ID: 2 / Number of copies: 6 / Enantiomer: LEVO / EC number: aspartate carbamoyltransferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 34.337105 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ANPLYQKHII SINDLSRDDL NLVLATAAKL KANPQPELLK HKVIASCFFE ASTRTRLSFE TSMHRLGASV VGFSDSANTS LGKKGETLA DTISVISTYV DAIVMRHPQE GAARLATEFS GNVPVLNAGD GSNQHPTQTL LDLFTIQETQ GRLDNLHVAM V GDLKYGRT ...String: ANPLYQKHII SINDLSRDDL NLVLATAAKL KANPQPELLK HKVIASCFFE ASTRTRLSFE TSMHRLGASV VGFSDSANTS LGKKGETLA DTISVISTYV DAIVMRHPQE GAARLATEFS GNVPVLNAGD GSNQHPTQTL LDLFTIQETQ GRLDNLHVAM V GDLKYGRT VHSLTQALAK FDGNRFYFIA PDALAMPQYI LDMLDEKGIA WSLHSSIEEV MAEVDILYMT RVQKERLDPS EY ANVKAQF VLRASDLHNA KANMKVLHPL PRVDEIATDV DKTPHAWYFQ QAGNGIFARQ ALLALVLNRD LVL UniProtKB: Aspartate carbamoyltransferase |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 6 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: CYTIDINE-5'-TRIPHOSPHATE
| Macromolecule | Name: CYTIDINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 12 / Formula: CTP |
|---|---|
| Molecular weight | Theoretical: 483.156 Da |
| Chemical component information |  ChemComp-CTP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 6 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: PHOSPHORIC ACID MONO(FORMAMIDE)ESTER
| Macromolecule | Name: PHOSPHORIC ACID MONO(FORMAMIDE)ESTER / type: ligand / ID: 6 / Number of copies: 6 / Formula: CP |
|---|---|
| Molecular weight | Theoretical: 141.02 Da |
| Chemical component information | 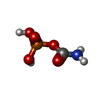 ChemComp-CP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE Details: Grids were manually coated with graphene-oxide following glow-discharge as is described in the associated publication. | ||||||||||||
| Vitrification | Cryogen name: ETHANE | ||||||||||||
| Details | E. coli aspartate transcarbamoylase was isolated in a neutral buffer and complexed with CP and nucleotide effector CTP in the presence of a physilogical concentration of Mg2+ |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 51.75 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































