[English] 日本語
 Yorodumi
Yorodumi- EMDB-4433: Thermophage P23-45 empty expanded capsid icosahedral reconstruction -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4433 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Thermophage P23-45 empty expanded capsid icosahedral reconstruction | |||||||||||||||||||||
 Map data Map data | Bacteriophage P23-45 empty expanded capsid icosahedral reconstruction | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | bacteriophage / thermophage / caudovirales / siphoviridae / capsid / auxiliary / HK97 / virus | |||||||||||||||||||||
| Function / homology | Major capsid protein GpE / Phage major capsid protein E / viral capsid, decoration / T=7 icosahedral viral capsid / host cell cytoplasm / Decoration protein / Major capsid protein Function and homology information Function and homology information | |||||||||||||||||||||
| Biological species |  Thermus virus P23-45 / Thermus virus P23-45 /   Thermus phage P2345 (virus) Thermus phage P2345 (virus) | |||||||||||||||||||||
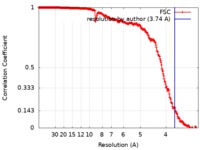
| Method | single particle reconstruction / cryo EM / Resolution: 3.74 Å | |||||||||||||||||||||
 Authors Authors | Bayfield OW / Klimuk E / Winkler DC / Hesketh EL / Chechik M / Cheng N / Dykeman EC / Minakhin L / Ranson NA / Severinov K ...Bayfield OW / Klimuk E / Winkler DC / Hesketh EL / Chechik M / Cheng N / Dykeman EC / Minakhin L / Ranson NA / Severinov K / Steven AC / Antson AA | |||||||||||||||||||||
| Funding support |  United Kingdom, United Kingdom,  United States, United States,  Russian Federation, 6 items Russian Federation, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: Cryo-EM structure and in vitro DNA packaging of a thermophilic virus with supersized T=7 capsids. Authors: Oliver W Bayfield / Evgeny Klimuk / Dennis C Winkler / Emma L Hesketh / Maria Chechik / Naiqian Cheng / Eric C Dykeman / Leonid Minakhin / Neil A Ranson / Konstantin Severinov / Alasdair C ...Authors: Oliver W Bayfield / Evgeny Klimuk / Dennis C Winkler / Emma L Hesketh / Maria Chechik / Naiqian Cheng / Eric C Dykeman / Leonid Minakhin / Neil A Ranson / Konstantin Severinov / Alasdair C Steven / Alfred A Antson /    Abstract: Double-stranded DNA viruses, including bacteriophages and herpesviruses, package their genomes into preformed capsids, using ATP-driven motors. Seeking to advance structural and mechanistic ...Double-stranded DNA viruses, including bacteriophages and herpesviruses, package their genomes into preformed capsids, using ATP-driven motors. Seeking to advance structural and mechanistic understanding, we established in vitro packaging for a thermostable bacteriophage, P23-45 of Both the unexpanded procapsid and the expanded mature capsid can package DNA in the presence of packaging ATPase over the 20 °C to 70 °C temperature range, with optimum activity at 50 °C to 65 °C. Cryo-EM reconstructions for the mature and immature capsids at 3.7-Å and 4.4-Å resolution, respectively, reveal conformational changes during capsid expansion. Capsomer interactions in the expanded capsid are reinforced by formation of intersubunit β-sheets with N-terminal segments of auxiliary protein trimers. Unexpectedly, the capsid has T=7 quasi-symmetry, despite the P23-45 genome being twice as large as those of known T=7 phages, in which the DNA is compacted to near-crystalline density. Our data explain this anomaly, showing how the canonical HK97 fold has adapted to double the volume of the capsid, while maintaining its structural integrity. Reconstructions of the procapsid and the expanded capsid defined the structure of the single vertex containing the portal protein. Together with a 1.95-Å resolution crystal structure of the portal protein and DNA packaging assays, these reconstructions indicate that capsid expansion affects the conformation of the portal protein, while still allowing DNA to be packaged. These observations suggest a mechanism by which structural events inside the capsid can be communicated to the outside. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4433.map.gz emd_4433.map.gz | 1.1 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4433-v30.xml emd-4433-v30.xml emd-4433.xml emd-4433.xml | 15.2 KB 15.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4433_fsc.xml emd_4433_fsc.xml | 24.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_4433.png emd_4433.png | 161.6 KB | ||
| Filedesc metadata |  emd-4433.cif.gz emd-4433.cif.gz | 6.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4433 http://ftp.pdbj.org/pub/emdb/structures/EMD-4433 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4433 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4433 | HTTPS FTP |
-Related structure data
| Related structure data |  6i9eMC  4445C  4446C  4447C  6ibcC  6ibgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4433.map.gz / Format: CCP4 / Size: 1.2 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4433.map.gz / Format: CCP4 / Size: 1.2 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Bacteriophage P23-45 empty expanded capsid icosahedral reconstruction | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.5975 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Thermus phage P2345
| Entire | Name:   Thermus phage P2345 (virus) Thermus phage P2345 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Thermus phage P2345
| Supramolecule | Name: Thermus phage P2345 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 466051 / Sci species name: Thermus phage P2345 / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:   Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) |
| Molecular weight | Theoretical: 26.57 MDa |
| Virus shell | Shell ID: 1 / T number (triangulation number): 7 |
-Macromolecule #1: Major head protein
| Macromolecule | Name: Major head protein / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermus virus P23-45 Thermus virus P23-45 |
| Molecular weight | Theoretical: 46.680754 KDa |
| Sequence | String: MRVPININNA LARVRDPLSI GGLKFPTTKE IQEAVAAIAD KFNQENDLVD RFFPEDSTFA SELELYLLRT QDAEQTGMTF VHQVGSTSL PVEARVAKVD LAKATWSPLA FKESRVWDEK EILYLGRLAD EVQAGVINEQ IAESLTWLMA RMRNRRRWLT W QVMRTGRI ...String: MRVPININNA LARVRDPLSI GGLKFPTTKE IQEAVAAIAD KFNQENDLVD RFFPEDSTFA SELELYLLRT QDAEQTGMTF VHQVGSTSL PVEARVAKVD LAKATWSPLA FKESRVWDEK EILYLGRLAD EVQAGVINEQ IAESLTWLMA RMRNRRRWLT W QVMRTGRI TIQPNDPYNP NGLKYVIDYG VTDIELPLPQ KFDAKDGNGN SAVDPIQYFR DLIKAATYFP DRRPVAIIVG PG FDEVLAD NTFVQKYVEY EKGWVVGQNT VQPPREVYRQ AALDIFKRYT GLEVMVYDKT YRDQDGSVKY WIPVGELIVL NQS TGPVGR FVYTAHVAGQ RNGKVVYATG PYLTVKDHLQ DDPPYYAIIA GFHGLPQLSG YNTEDFSFHR FKWLKYANNV QSYL PPFPP KVEL UniProtKB: Major capsid protein |
-Macromolecule #2: Auxiliary protein
| Macromolecule | Name: Auxiliary protein / type: protein_or_peptide / ID: 2 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermus virus P23-45 Thermus virus P23-45 |
| Molecular weight | Theoretical: 16.661002 KDa |
| Sequence | String: MDKVKLFQTI GRVEYWERVP RLHAYGVFAL PFPMDPDVNW AQWFTGPHPR AFLVSIHKYG PKAGHVYPTN LTDEDALLNV IGMVLDGHD YENDPNVTVT LKAAVPIEYV QQDPQAPALQ PHQAVLDAAE VLKLKVIKGH YFFDYTR UniProtKB: Decoration protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: AIR | ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Average exposure time: 2.0 sec. / Average electron dose: 99.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 75000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6i9e: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)