+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human Wnt3a bound to WLS and CALR | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | SIGNALING PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationWnt signaling pathway involved in forebrain neuroblast division / positive regulation of dermatome development / calcium ion transmembrane transport via low voltage-gated calcium channel / positive regulation of collateral sprouting in absence of injury / positive regulation of mesodermal cell fate specification / paraxial mesodermal cell fate commitment / axis elongation involved in somitogenesis / cell proliferation in midbrain / spinal cord association neuron differentiation / Wnt protein secretion ...Wnt signaling pathway involved in forebrain neuroblast division / positive regulation of dermatome development / calcium ion transmembrane transport via low voltage-gated calcium channel / positive regulation of collateral sprouting in absence of injury / positive regulation of mesodermal cell fate specification / paraxial mesodermal cell fate commitment / axis elongation involved in somitogenesis / cell proliferation in midbrain / spinal cord association neuron differentiation / Wnt protein secretion / Formation of the posterior neural plate / response to biphenyl / Calnexin/calreticulin cycle / COP9 signalosome assembly / cytolytic granule / Wnt-Frizzled-LRP5/6 complex / positive regulation of Wnt protein secretion / Negative regulation of TCF-dependent signaling by WNT ligand antagonists / positive regulation of cell-cell adhesion mediated by cadherin / positive regulation of dendritic cell chemotaxis / WNT ligand biogenesis and trafficking / Signaling by RNF43 mutants / nuclear receptor-mediated glucocorticoid signaling pathway / negative regulation of axon extension involved in axon guidance / Assembly of Viral Components at the Budding Site / ATF6 (ATF6-alpha) activates chaperone genes / synaptic vesicle recycling / cell proliferation in forebrain / negative regulation of trophoblast cell migration / positive regulation of cardiac muscle cell differentiation / cortical granule / cellular response to electrical stimulus / response to peptide / Specification of the neural plate border / regulation of meiotic nuclear division / complement component C1q complex binding / sequestering of calcium ion / cementum mineralization / negative regulation of retinoic acid receptor signaling pathway / secondary palate development / presynapse assembly / endoplasmic reticulum quality control compartment / somatic stem cell division / protein folding in endoplasmic reticulum / sarcoplasmic reticulum lumen / cardiac muscle cell fate commitment / non-canonical Wnt signaling pathway / co-receptor binding / hindbrain development / positive regulation of skeletal muscle tissue development / negative regulation of dopaminergic neuron differentiation / Wnt-protein binding / negative regulation of intracellular steroid hormone receptor signaling pathway / nuclear export signal receptor activity / regulation of postsynapse to nucleus signaling pathway / dorsal/ventral neural tube patterning / cardiac muscle cell differentiation / post-anal tail morphogenesis / positive regulation of hepatocyte proliferation / midbrain dopaminergic neuron differentiation / mammary gland development / frizzled binding / positive regulation of neural precursor cell proliferation / exocrine pancreas development / myoblast differentiation / Class B/2 (Secretin family receptors) / Disassembly of the destruction complex and recruitment of AXIN to the membrane / anterior/posterior axis specification / cortical actin cytoskeleton organization / response to glycoside / Scavenging by Class A Receptors / inner ear morphogenesis / nuclear androgen receptor binding / Scavenging by Class F Receptors / cellular response to lithium ion / negative regulation of neuron differentiation / negative regulation of fat cell differentiation / Formation of paraxial mesoderm / heart looping / response to testosterone / regulation of synapse organization / midbrain development / organelle membrane / fat cell differentiation / smooth endoplasmic reticulum / hormone binding / B cell proliferation / mesoderm formation / hemopoiesis / positive regulation of receptor internalization / skeletal muscle cell differentiation / molecular sequestering activity / protein localization to nucleus / cell fate commitment / positive regulation of Wnt signaling pathway / canonical Wnt signaling pathway / regulation of presynapse assembly / positive regulation of cell cycle / cellular response to retinoic acid / positive regulation of B cell proliferation Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||
 Authors Authors | Qi X / Hu Q / Li X | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Molecular basis of Wnt biogenesis, secretion, and Wnt7-specific signaling. Authors: Xiaofeng Qi / Qinli Hu / Nadia Elghobashi-Meinhardt / Tao Long / Hongwen Chen / Xiaochun Li /   Abstract: Wnt proteins are enzymatically lipidated by Porcupine (PORCN) in the ER and bind to Wntless (WLS) for intracellular transport and secretion. Mechanisms governing the transfer of these low-solubility ...Wnt proteins are enzymatically lipidated by Porcupine (PORCN) in the ER and bind to Wntless (WLS) for intracellular transport and secretion. Mechanisms governing the transfer of these low-solubility Wnts from the ER to the extracellular space remain unclear. Through structural and functional analyses of Wnt7a, a crucial Wnt involved in central nervous system angiogenesis and blood-brain barrier maintenance, we have elucidated the principles of Wnt biogenesis and Wnt7-specific signaling. The Wnt7a-WLS complex binds to calreticulin (CALR), revealing that CALR functions as a chaperone to facilitate Wnt transfer from PORCN to WLS during Wnt biogenesis. Our structures, functional analyses, and molecular dynamics simulations demonstrate that a phospholipid in the core of Wnt-bound WLS regulates the association and dissociation between Wnt and WLS, suggesting a lipid-mediated Wnt secretion mechanism. Finally, the structure of Wnt7a bound to RECK, a cell-surface Wnt7 co-receptor, reveals how RECK engages the N-terminal domain of Wnt7a to activate Wnt7-specific signaling. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41767.map.gz emd_41767.map.gz | 78.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41767-v30.xml emd-41767-v30.xml emd-41767.xml emd-41767.xml | 20.5 KB 20.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_41767.png emd_41767.png | 55.2 KB | ||
| Filedesc metadata |  emd-41767.cif.gz emd-41767.cif.gz | 7.1 KB | ||
| Others |  emd_41767_half_map_1.map.gz emd_41767_half_map_1.map.gz emd_41767_half_map_2.map.gz emd_41767_half_map_2.map.gz | 77.6 MB 77.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41767 http://ftp.pdbj.org/pub/emdb/structures/EMD-41767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41767 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41767 | HTTPS FTP |
-Validation report
| Summary document |  emd_41767_validation.pdf.gz emd_41767_validation.pdf.gz | 941.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41767_full_validation.pdf.gz emd_41767_full_validation.pdf.gz | 941.2 KB | Display | |
| Data in XML |  emd_41767_validation.xml.gz emd_41767_validation.xml.gz | 12.9 KB | Display | |
| Data in CIF |  emd_41767_validation.cif.gz emd_41767_validation.cif.gz | 15.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41767 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41767 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41767 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41767 | HTTPS FTP |
-Related structure data
| Related structure data |  8tzrMC  8tzoC  8tzpC  8tzsC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41767.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41767.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41767_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_41767_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Wnt3a-WLS-CALR Complex
| Entire | Name: Wnt3a-WLS-CALR Complex |
|---|---|
| Components |
|
-Supramolecule #1: Wnt3a-WLS-CALR Complex
| Supramolecule | Name: Wnt3a-WLS-CALR Complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Protein Wnt-3a
| Macromolecule | Name: Protein Wnt-3a / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.421832 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAPLGYFLLL CSLKQALGSY PIWWSLAVGP QYSSLGSQPI LCASIPGLVP KQLRFCRNYV EIMPSVAEGI KIGIQECQHQ FRGRRWNCT TVHDSLAIFG PVLDKATRES AFVHAIASAG VAFAVTRSCA EGTAAICGCS SRHQGSPGKG WKWGGCSEDI E FGGMVSRE ...String: MAPLGYFLLL CSLKQALGSY PIWWSLAVGP QYSSLGSQPI LCASIPGLVP KQLRFCRNYV EIMPSVAEGI KIGIQECQHQ FRGRRWNCT TVHDSLAIFG PVLDKATRES AFVHAIASAG VAFAVTRSCA EGTAAICGCS SRHQGSPGKG WKWGGCSEDI E FGGMVSRE FADARENRPD ARSAMNRHNN EAGRQAIASH MHLKCKCHGL SGSCEVKTCW WSQPDFRAIG DFLKDKYDSA SE MVVEKHR ESRGWVETLR PRYTYFKVPT ERDLVYYEAS PNFCEPNPET GSFGTRDRTC NVSSHGIDGC DLLCCGRGHN ARA ERRREK CRCVFHWCCY VSCQECTRVY DVHTCK UniProtKB: Protein Wnt-3a |
-Macromolecule #2: Protein wntless homolog
| Macromolecule | Name: Protein wntless homolog / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 62.317973 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAGAIIENMS TKKLCIVGGI LLVFQIIAFL VGGLIAPGPT TAVSYMSVKC VDARKNHHKT KWFVPWGPNH CDKIRDIEEA IPREIEAND IVFSVHIPLP HMEMSPWFQF MLFILQLDIA FKLNNQIREN AEVSMDVSLA YRDDAFAEWT EMAHERVPRK L KCTFTSPK ...String: MAGAIIENMS TKKLCIVGGI LLVFQIIAFL VGGLIAPGPT TAVSYMSVKC VDARKNHHKT KWFVPWGPNH CDKIRDIEEA IPREIEAND IVFSVHIPLP HMEMSPWFQF MLFILQLDIA FKLNNQIREN AEVSMDVSLA YRDDAFAEWT EMAHERVPRK L KCTFTSPK TPEHEGRYYE CDVLPFMEIG SVAHKFYLLN IRLPVNEKKK INVGIGEIKD IRLVGIHQNG GFTKVWFAMK TF LTPSIFI IMVWYWRRIT MMSRPPVLLE KVIFALGISM TFINIPVEWF SIGFDWTWML LFGDIRQGIF YAMLLSFWII FCG EHMMDQ HERNHIAGYW KQVGPIAVGS FCLFIFDMCE RGVQLTNPFY SIWTTDIGTE LAMAFIIVAG ICLCLYFLFL CFMV FQVFR NISGKQSSLP AMSKVRRLHY EGLIFRFKFL MLITLACAAM TVIFFIVSQV TEGHWKWGGV TVQVNSAFFT GIYGM WNLY VFALMFLYAP SHKNYGEDQS NGDLGVHSGE ELQLTTTITH VDGPTEIYKL TRKEAQE UniProtKB: Protein wntless homolog |
-Macromolecule #3: Calreticulin
| Macromolecule | Name: Calreticulin / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 48.198379 KDa |
| Sequence | String: MLLSVPLLLG LLGLAVAEPA VYFKEQFLDG DGWTSRWIES KHKSDFGKFV LSSGKFYGDE EKDKGLQTSQ DARFYALSAS FEPFSNKGQ TLVVQFTVKH EQNIDCGGGY VKLFPNSLDQ TDMHGDSEYN IMFGPDICGP GTKKVHVIFN YKGKNVLINK D IRCKDDEF ...String: MLLSVPLLLG LLGLAVAEPA VYFKEQFLDG DGWTSRWIES KHKSDFGKFV LSSGKFYGDE EKDKGLQTSQ DARFYALSAS FEPFSNKGQ TLVVQFTVKH EQNIDCGGGY VKLFPNSLDQ TDMHGDSEYN IMFGPDICGP GTKKVHVIFN YKGKNVLINK D IRCKDDEF THLYTLIVRP DNTYEVKIDN SQVESGSLED DWDFLPPKKI KDPDASKPED WDERAKIDDP TDSKPEDWDK PE HIPDPDA KKPEDWDEEM DGEWEPPVIQ NPEYKGEWKP RQIDNPDYKG TWIHPEIDNP EYSPDPSIYA YDNFGVLGLD LWQ VKSGTI FDNFLITNDE AYAEEFGNET WGVTKAAEKQ MKDKQDEEQR LKEEEEDKKR KEEEEAEDKE DDEDKDEDEE DEED KEEDE EEDVPGQAKD EL UniProtKB: Calreticulin |
-Macromolecule #5: PALMITOLEIC ACID
| Macromolecule | Name: PALMITOLEIC ACID / type: ligand / ID: 5 / Number of copies: 1 / Formula: PAM |
|---|---|
| Molecular weight | Theoretical: 254.408 Da |
| Chemical component information | 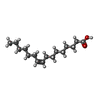 ChemComp-PAM: |
-Macromolecule #6: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 6 / Number of copies: 1 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)