[English] 日本語
 Yorodumi
Yorodumi- EMDB-4132: Structure of the mammalian ribosomal termination complex with acc... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4132 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the mammalian ribosomal termination complex with accommodated eRF1 | ||||||||||||
 Map data Map data | Postprocessed, sharpened map. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Translation / Elongation / Ribosome | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationtranslation termination factor activity / translation release factor complex / cytoplasmic translational termination / regulation of translational termination / translation release factor activity / translation release factor activity, codon specific / protein methylation / sequence-specific mRNA binding / peptidyl-tRNA hydrolase activity / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay ...translation termination factor activity / translation release factor complex / cytoplasmic translational termination / regulation of translational termination / translation release factor activity / translation release factor activity, codon specific / protein methylation / sequence-specific mRNA binding / peptidyl-tRNA hydrolase activity / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / Protein hydroxylation / Eukaryotic Translation Termination / 90S preribosome / ubiquitin ligase inhibitor activity / positive regulation of signal transduction by p53 class mediator / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / translational termination / phagocytic cup / protein-RNA complex assembly / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / translation regulator activity / rough endoplasmic reticulum / ribosomal small subunit export from nucleus / gastrulation / MDM2/MDM4 family protein binding / cytosolic ribosome / class I DNA-(apurinic or apyrimidinic site) endonuclease activity / DNA-(apurinic or apyrimidinic site) lyase / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / positive regulation of apoptotic signaling pathway / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA / small-subunit processome / spindle / Regulation of expression of SLITs and ROBOs / rRNA processing / positive regulation of canonical Wnt signaling pathway / regulation of translation / rhythmic process / antimicrobial humoral immune response mediated by antimicrobial peptide / large ribosomal subunit / ribosomal small subunit assembly / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / killing of cells of another organism / cytosolic large ribosomal subunit / defense response to Gram-negative bacterium / perikaryon / cell differentiation / cytoplasmic translation / tRNA binding / mitochondrial inner membrane / postsynaptic density / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / cell division / DNA repair / mRNA binding / apoptotic process / centrosome / synapse / dendrite / nucleolus / perinuclear region of cytoplasm / endoplasmic reticulum / Golgi apparatus / DNA binding / RNA binding / zinc ion binding / membrane / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
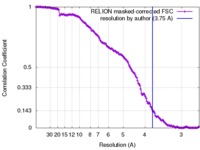
| Method | single particle reconstruction / cryo EM / Resolution: 3.75 Å | ||||||||||||
 Authors Authors | Shao S / Murray J | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell / Year: 2016 Journal: Cell / Year: 2016Title: Decoding Mammalian Ribosome-mRNA States by Translational GTPase Complexes. Authors: Sichen Shao / Jason Murray / Alan Brown / Jack Taunton / V Ramakrishnan / Ramanujan S Hegde /   Abstract: In eukaryotes, accurate protein synthesis relies on a family of translational GTPases that pair with specific decoding factors to decipher the mRNA code on ribosomes. We present structures of the ...In eukaryotes, accurate protein synthesis relies on a family of translational GTPases that pair with specific decoding factors to decipher the mRNA code on ribosomes. We present structures of the mammalian ribosome engaged with decoding factor⋅GTPase complexes representing intermediates of translation elongation (aminoacyl-tRNA⋅eEF1A), termination (eRF1⋅eRF3), and ribosome rescue (Pelota⋅Hbs1l). Comparative analyses reveal that each decoding factor exploits the plasticity of the ribosomal decoding center to differentially remodel ribosomal proteins and rRNA. This leads to varying degrees of large-scale ribosome movements and implies distinct mechanisms for communicating information from the decoding center to each GTPase. Additional structural snapshots of the translation termination pathway reveal the conformational changes that choreograph the accommodation of decoding factors into the peptidyl transferase center. Our results provide a structural framework for how different states of the mammalian ribosome are selectively recognized by the appropriate decoding factor⋅GTPase complex to ensure translational fidelity. #1:  Journal: To Be Published Journal: To Be PublishedTitle: Decoding mammalian ribosome-mRNA states by translational GTPase complexes Authors: Shao S / Murray J / Brown A / Taunton J / Ramakrishnan V / Hegde RS | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4132.map.gz emd_4132.map.gz | 14.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4132-v30.xml emd-4132-v30.xml emd-4132.xml emd-4132.xml | 106.1 KB 106.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4132_fsc.xml emd_4132_fsc.xml | 14.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_4132.png emd_4132.png | 192.8 KB | ||
| Filedesc metadata |  emd-4132.cif.gz emd-4132.cif.gz | 20.9 KB | ||
| Others |  emd_4132_half_map_1.map.gz emd_4132_half_map_1.map.gz emd_4132_half_map_2.map.gz emd_4132_half_map_2.map.gz | 250.3 MB 250.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4132 http://ftp.pdbj.org/pub/emdb/structures/EMD-4132 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4132 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4132 | HTTPS FTP |
-Related structure data
| Related structure data |  5lzuMC  4129C  4130C  4131C  4133C  4134C  4135C  4136C  4137C  5lzsC  5lztC  5lzvC  5lzwC  5lzxC  5lzyC  5lzzC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4132.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4132.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed, sharpened map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Half map 1.
| File | emd_4132_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2.
| File | emd_4132_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Affinity-purified 80S ribosome-nascent chain complex reconstitute...
+Supramolecule #1: Affinity-purified 80S ribosome-nascent chain complex reconstitute...
+Macromolecule #1: uL2
+Macromolecule #2: uL3
+Macromolecule #3: uL4
+Macromolecule #4: 60S ribosomal protein L5
+Macromolecule #5: 60S ribosomal protein L6
+Macromolecule #6: uL30
+Macromolecule #7: eL8
+Macromolecule #8: uL6
+Macromolecule #9: uL16
+Macromolecule #10: uL5
+Macromolecule #11: eL13
+Macromolecule #12: eL14
+Macromolecule #13: Ribosomal protein L15
+Macromolecule #14: uL13
+Macromolecule #15: uL22
+Macromolecule #16: eL18
+Macromolecule #17: eL19
+Macromolecule #18: eL20
+Macromolecule #19: eL21
+Macromolecule #20: eL22
+Macromolecule #21: uL14
+Macromolecule #22: eL24
+Macromolecule #23: uL23
+Macromolecule #24: uL24
+Macromolecule #25: 60S ribosomal protein L27
+Macromolecule #26: uL15
+Macromolecule #27: eL29
+Macromolecule #28: eL30
+Macromolecule #29: eL31
+Macromolecule #30: eL32
+Macromolecule #31: eL33
+Macromolecule #32: eL34
+Macromolecule #33: uL29
+Macromolecule #34: 60S ribosomal protein L36
+Macromolecule #35: Ribosomal protein L37
+Macromolecule #36: eL38
+Macromolecule #37: eL39
+Macromolecule #38: eL40
+Macromolecule #39: 60s ribosomal protein l41
+Macromolecule #40: eL42
+Macromolecule #41: eL43
+Macromolecule #42: eL28
+Macromolecule #43: uL10
+Macromolecule #44: uL11
+Macromolecule #51: uS2
+Macromolecule #52: 40S ribosomal protein S3a
+Macromolecule #53: uS5
+Macromolecule #54: uS3
+Macromolecule #55: 40S ribosomal protein S4
+Macromolecule #56: uS7
+Macromolecule #57: 40S ribosomal protein S6
+Macromolecule #58: eS7
+Macromolecule #59: 40S ribosomal protein S8
+Macromolecule #60: Ribosomal protein S9 (Predicted)
+Macromolecule #61: eS10
+Macromolecule #62: uS17
+Macromolecule #63: 40S ribosomal protein S12
+Macromolecule #64: uS15
+Macromolecule #65: uS11
+Macromolecule #66: uS19
+Macromolecule #67: uS9
+Macromolecule #68: eS17
+Macromolecule #69: uS13
+Macromolecule #70: eS19
+Macromolecule #71: uS10
+Macromolecule #72: eS21
+Macromolecule #73: uS8
+Macromolecule #74: uS12
+Macromolecule #75: eS24
+Macromolecule #76: eS25
+Macromolecule #77: eS26
+Macromolecule #78: 40S ribosomal protein S27
+Macromolecule #79: eS28
+Macromolecule #80: uS14
+Macromolecule #81: eS30
+Macromolecule #82: eS31
+Macromolecule #83: RACK1
+Macromolecule #85: Eukaryotic peptide chain release factor subunit 1
+Macromolecule #45: P-site tRNA
+Macromolecule #46: E-site tRNA
+Macromolecule #47: 28S ribosomal RNA
+Macromolecule #48: 5S ribosomal RNA
+Macromolecule #49: 5.8S ribosomal RNA
+Macromolecule #50: 18S ribosomal RNA
+Macromolecule #84: mRNA (UGA stop codon)
+Macromolecule #86: MAGNESIUM ION
+Macromolecule #87: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 5 / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III Details: 3 ul aliquots were applied to the grid and incubated for 30 s, before blotting for 3s to remove excess solution.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Number real images: 1611 / Average exposure time: 1.0 sec. / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated magnification: 104478 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Overall B value: 63.8 / Target criteria: FSCaverage |
|---|---|
| Output model |  PDB-5lzu: |
 Movie
Movie Controller
Controller
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)