+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3687 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a pre-catalytic spliceosome (B6 map) | |||||||||
 Map data Map data | sharpened B6 map | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | |||||||||
 Authors Authors | Plaschka C / Lin P-C / Nagai K | |||||||||
 Citation Citation |  Journal: Nature / Year: 2017 Journal: Nature / Year: 2017Title: Structure of a pre-catalytic spliceosome. Authors: Clemens Plaschka / Pei-Chun Lin / Kiyoshi Nagai /  Abstract: Intron removal requires assembly of the spliceosome on precursor mRNA (pre-mRNA) and extensive remodelling to form the spliceosome's catalytic centre. Here we report the cryo-electron microscopy ...Intron removal requires assembly of the spliceosome on precursor mRNA (pre-mRNA) and extensive remodelling to form the spliceosome's catalytic centre. Here we report the cryo-electron microscopy structure of the yeast Saccharomyces cerevisiae pre-catalytic B complex spliceosome at near-atomic resolution. The mobile U2 small nuclear ribonucleoprotein particle (snRNP) associates with U4/U6.U5 tri-snRNP through the U2/U6 helix II and an interface between U4/U6 di-snRNP and the U2 snRNP SF3b-containing domain, which also transiently contacts the helicase Brr2. The 3' region of the U2 snRNP is flexibly attached to the SF3b-containing domain and protrudes over the concave surface of tri-snRNP, where the U1 snRNP may reside before its release from the pre-mRNA 5' splice site. The U6 ACAGAGA sequence forms a hairpin that weakly tethers the 5' splice site. The B complex proteins Prp38, Snu23 and Spp381 bind the Prp8 N-terminal domain and stabilize U6 ACAGAGA stem-pre-mRNA and Brr2-U4 small nuclear RNA interactions. These results provide important insights into the events leading to active site formation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3687.map.gz emd_3687.map.gz | 389.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3687-v30.xml emd-3687-v30.xml emd-3687.xml emd-3687.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3687.png emd_3687.png | 13.2 KB | ||
| Others |  emd_3687_additional.map.gz emd_3687_additional.map.gz | 379.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3687 http://ftp.pdbj.org/pub/emdb/structures/EMD-3687 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3687 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3687 | HTTPS FTP |
-Related structure data
| Related structure data |  3682C  3683C  3684C  3685C  3686C  3688C  5nrlC C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10180 (Title: Structure of a pre-catalytic spliceosome / Data size: 126.5 EMPIAR-10180 (Title: Structure of a pre-catalytic spliceosome / Data size: 126.5 Data #1: Polished parcitles [picked particles - single frame - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3687.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3687.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened B6 map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.43 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
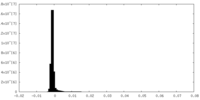
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unsharpened B6 map
| File | emd_3687_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened B6 map | ||||||||||||
| Projections & Slices |
| ||||||||||||
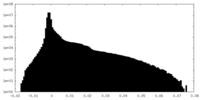
| Density Histograms |
- Sample components
Sample components
-Entire : Pre-catalytic B complex Spliceosome
| Entire | Name: Pre-catalytic B complex Spliceosome |
|---|---|
| Components |
|
-Supramolecule #1: Pre-catalytic B complex Spliceosome
| Supramolecule | Name: Pre-catalytic B complex Spliceosome / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 2.5 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.9 Component:
Details: Buffer pH: HEPES, 7.9; EDTA, 8.0 | ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK III Details: Grids were glow-discharged for 15 s before deposition of 3 microliter sample (~1.5 mg mL-1), and subsequently incubated for 2-3.5 s before blotting and vitrification by plunging into liquid ...Details: Grids were glow-discharged for 15 s before deposition of 3 microliter sample (~1.5 mg mL-1), and subsequently incubated for 2-3.5 s before blotting and vitrification by plunging into liquid ethane with a Vitrobot Mark III (FEI) operated at 4 degrees Celsius and 100% humidity.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Sampling interval: 5.0 µm / Digitization - Frames/image: 1-20 / Number real images: 5115 / Average exposure time: 16.0 sec. / Average electron dose: 56.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 5.3 µm / Calibrated defocus min: 0.35 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Overall B value: 140 |
|---|
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)