+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Pseudomonas phage E217 baseplate complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Pseudomonas / phage / baseplate / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology information: / : / Baseplate hub gp41 / Baseplate component gp38 / Protein of unknown function DUF2612 / : / Bacteriophage baseplate wedge 1 protein / Tail fiber protein gp32 / : / Cyanophage baseplate Pam3 plug gp18 ...: / : / Baseplate hub gp41 / Baseplate component gp38 / Protein of unknown function DUF2612 / : / Bacteriophage baseplate wedge 1 protein / Tail fiber protein gp32 / : / Cyanophage baseplate Pam3 plug gp18 / Phage protein Gp138 N-terminal domain / Phage protein Gp138 N-terminal domain / Protein of unknown function DUF2634 / Contractile injection system sheath initiator / Baseplate protein J-like / Baseplate J-like protein barrel domain / Vgr protein, OB-fold domain superfamily Similarity search - Domain/homology | |||||||||
| Biological species |  Pseudomonas phage vB_PaeM_E217 (virus) Pseudomonas phage vB_PaeM_E217 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Li F / Cingolani G / Hou C | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: High-resolution cryo-EM structure of the Pseudomonas bacteriophage E217. Authors: Fenglin Li / Chun-Feng David Hou / Ravi K Lokareddy / Ruoyu Yang / Francesca Forti / Federica Briani / Gino Cingolani /   Abstract: E217 is a Pseudomonas phage used in an experimental cocktail to eradicate cystic fibrosis-associated Pseudomonas aeruginosa. Here, we describe the structure of the whole E217 virion before and after ...E217 is a Pseudomonas phage used in an experimental cocktail to eradicate cystic fibrosis-associated Pseudomonas aeruginosa. Here, we describe the structure of the whole E217 virion before and after DNA ejection at 3.1 Å and 4.5 Å resolution, respectively, determined using cryogenic electron microscopy (cryo-EM). We identify and build de novo structures for 19 unique E217 gene products, resolve the tail genome-ejection machine in both extended and contracted states, and decipher the complete architecture of the baseplate formed by 66 polypeptide chains. We also determine that E217 recognizes the host O-antigen as a receptor, and we resolve the N-terminal portion of the O-antigen-binding tail fiber. We propose that E217 design principles presented in this paper are conserved across PB1-like Myoviridae phages of the Pbunavirus genus that encode a ~1.4 MDa baseplate, dramatically smaller than the coliphage T4. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28405.map.gz emd_28405.map.gz | 225.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28405-v30.xml emd-28405-v30.xml emd-28405.xml emd-28405.xml | 24.5 KB 24.5 KB | Display Display |  EMDB header EMDB header |
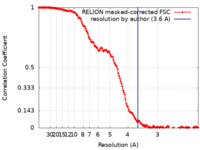
| FSC (resolution estimation) |  emd_28405_fsc.xml emd_28405_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_28405.png emd_28405.png | 107.9 KB | ||
| Filedesc metadata |  emd-28405.cif.gz emd-28405.cif.gz | 7.1 KB | ||
| Others |  emd_28405_half_map_1.map.gz emd_28405_half_map_1.map.gz emd_28405_half_map_2.map.gz emd_28405_half_map_2.map.gz | 193.5 MB 193.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28405 http://ftp.pdbj.org/pub/emdb/structures/EMD-28405 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28405 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28405 | HTTPS FTP |
-Related structure data
| Related structure data |  8eonMC  8envC  8frsC  8fuvC  8fvgC  8fvhC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28405.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28405.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_28405_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_28405_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Pseudomonas phage vB_PaeM_E217
+Supramolecule #1: Pseudomonas phage vB_PaeM_E217
+Macromolecule #1: Baseplate component gp33
+Macromolecule #2: Baseplate component gp34
+Macromolecule #3: Baseplate component gp36
+Macromolecule #4: Triplex gp44-b
+Macromolecule #5: Triplex gp45
+Macromolecule #6: Baseplate component gp37
+Macromolecule #7: Baseplate component gp38
+Macromolecule #8: Baseplate hub gp41
+Macromolecule #9: Baseplate spike gp43
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 1.25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)