+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2402 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 3D structure of a properdin vertex | |||||||||
 Map data Map data | Reconstruction of a vertex from a Properdin oligomer (tetramer) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | properdin / C3b / AP C3 convertase / complement / electron microscopy / EM | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoplasmic side of Golgi membrane / positive regulation of opsonization / Defective B3GALTL causes PpS / O-glycosylation of TSR domain-containing proteins / regulation of complement activation / Alternative complement activation / Activation of C3 and C5 / complement activation / complement activation, alternative pathway / Regulation of Complement cascade ...cytoplasmic side of Golgi membrane / positive regulation of opsonization / Defective B3GALTL causes PpS / O-glycosylation of TSR domain-containing proteins / regulation of complement activation / Alternative complement activation / Activation of C3 and C5 / complement activation / complement activation, alternative pathway / Regulation of Complement cascade / positive regulation of immune response / specific granule lumen / tertiary granule lumen / defense response to bacterium / immune response / endoplasmic reticulum lumen / Neutrophil degranulation / extracellular space / extracellular region Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 23.4 Å | |||||||||
 Authors Authors | Alcorlo M / Tortajada A / Rodriguez de Cordoba S / Llorca O | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2013 Journal: Proc Natl Acad Sci U S A / Year: 2013Title: Structural basis for the stabilization of the complement alternative pathway C3 convertase by properdin. Authors: Martín Alcorlo / Agustín Tortajada / Santiago Rodríguez de Córdoba / Oscar Llorca /  Abstract: Complement is an essential component of innate immunity. Its activation results in the assembly of unstable protease complexes, denominated C3/C5 convertases, leading to inflammation and lysis. ...Complement is an essential component of innate immunity. Its activation results in the assembly of unstable protease complexes, denominated C3/C5 convertases, leading to inflammation and lysis. Regulatory proteins inactivate C3/C5 convertases on host surfaces to avoid collateral tissue damage. On pathogen surfaces, properdin stabilizes C3/C5 convertases to efficiently fight infection. How properdin performs this function is, however, unclear. Using electron microscopy we show that the N- and C-terminal ends of adjacent monomers in properdin oligomers conform a curly vertex that holds together the AP convertase, interacting with both the C345C and vWA domains of C3b and Bb, respectively. Properdin also promotes a large displacement of the TED (thioester-containing domain) and CUB (complement protein subcomponents C1r/C1s, urchin embryonic growth factor and bone morphogenetic protein 1) domains of C3b, which likely impairs C3-convertase inactivation by regulatory proteins. The combined effect of molecular cross-linking and structural reorganization increases stability of the C3 convertase and facilitates recruitment of fluid-phase C3 convertase to the cell surfaces. Our model explains how properdin mediates the assembly of stabilized C3/C5-convertase clusters, which helps to localize complement amplification to pathogen surfaces. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2402.map.gz emd_2402.map.gz | 514.9 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2402-v30.xml emd-2402-v30.xml emd-2402.xml emd-2402.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2402.jpg emd_2402.jpg | 77.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2402 http://ftp.pdbj.org/pub/emdb/structures/EMD-2402 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2402 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2402 | HTTPS FTP |
-Validation report
| Summary document |  emd_2402_validation.pdf.gz emd_2402_validation.pdf.gz | 198.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2402_full_validation.pdf.gz emd_2402_full_validation.pdf.gz | 197.3 KB | Display | |
| Data in XML |  emd_2402_validation.xml.gz emd_2402_validation.xml.gz | 5.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2402 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2402 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2402 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2402 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2402.map.gz / Format: CCP4 / Size: 1.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2402.map.gz / Format: CCP4 / Size: 1.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of a vertex from a Properdin oligomer (tetramer) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.84 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
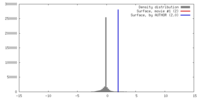
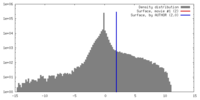
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human Properdin purified from plasma
| Entire | Name: Human Properdin purified from plasma |
|---|---|
| Components |
|
-Supramolecule #1000: Human Properdin purified from plasma
| Supramolecule | Name: Human Properdin purified from plasma / type: sample / ID: 1000 Details: The structure corresponds to a head-to-tail connection between two Properdin monomers (from a tetrameric species) Oligomeric state: dimer / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 45 KDa |
-Macromolecule #1: Complement Factor P
| Macromolecule | Name: Complement Factor P / type: protein_or_peptide / ID: 1 / Name.synonym: Properdin / Number of copies: 2 / Oligomeric state: dimer / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Tissue: plasma Homo sapiens (human) / synonym: Human / Tissue: plasma |
| Molecular weight | Experimental: 54 KDa / Theoretical: 53 KDa |
| Sequence | UniProtKB: Properdin GO: extracellular space, complement activation, alternative pathway, defense response to bacterium, regulation of complement activation InterPro: Thrombospondin type-1 (TSP1) repeat |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.01 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 20 mM Hepes, 75 mM NaCl and 5 mM MgCl2 |
| Staining | Type: NEGATIVE Details: Grids with adsorbed protein floated on 1% w/v uranyl acetate for 15 seconds |
| Grid | Details: 400 mesh copper carbon only (50ct), glow discharged |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 1230 |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected using a CMOS camera and the power spectrum |
| Details | Micrographs were recorded using a low-dose protocol under control of the EM-TOOLS software (TVIPS) |
| Date | Mar 21, 2012 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Digitization - Sampling interval: 2.84 µm / Number real images: 625 / Average electron dose: 10 e/Å2 / Od range: 1.4 / Bits/pixel: 16 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 100 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Calibrated magnification: 54926 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.9 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 40000 |
| Sample stage | Specimen holder model: JEOL / Tilt angle max: 40 |
- Image processing
Image processing
| Details | The particles were manually selected using Boxer (EMAN1). Images were classified and averaged using maximum-likelihood multi-reference methods as implemented in XMIPP. Ab initio templates for angular refinement were obtained using the command e2initial model in EMAN2. 3D reconstructions were obtained using angular refinement using EMAN. |
|---|---|
| CTF correction | Details: Each CCD Frame, estimated with CTFFIND and corrected using BSOFT |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 23.4 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN1, EMAN2, Xmipp-2.4 / Number images used: 6425 |
| Final two d classification | Number classes: 30 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  UCSF Chimera UCSF Chimera |
| Details | The domains were separately fitted using UCSF Chimera |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross correlation |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)