+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-2361 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Electron cryo-EM of the full-length Thermus thermophilus DNA gyrase in complex with a 155bp DNA and ciprofloxacin | |||||||||
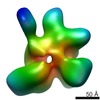
 マップデータ マップデータ | Recontruction of the full length Thermus thermophilus DNA gyrase bound to DNA. | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | DNA gyrase / DNA topoisomerase / DNA supercoiling | |||||||||
| 生物種 |   Thermus thermophilus (バクテリア) / Thermus thermophilus (バクテリア) /  | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 23.0 Å | |||||||||
 データ登録者 データ登録者 | Papillon J / Menetret JF / Batisse C / Helye R / Schultz P / Potier N / Lamour V | |||||||||
 引用 引用 |  ジャーナル: Nucleic Acids Res / 年: 2013 ジャーナル: Nucleic Acids Res / 年: 2013タイトル: Structural insight into negative DNA supercoiling by DNA gyrase, a bacterial type 2A DNA topoisomerase. 著者: Julie Papillon / Jean-François Ménétret / Claire Batisse / Reynald Hélye / Patrick Schultz / Noëlle Potier / Valérie Lamour /  要旨: Type 2A DNA topoisomerases (Topo2A) remodel DNA topology during replication, transcription and chromosome segregation. These multisubunit enzymes catalyze the transport of a double-stranded DNA ...Type 2A DNA topoisomerases (Topo2A) remodel DNA topology during replication, transcription and chromosome segregation. These multisubunit enzymes catalyze the transport of a double-stranded DNA through a transient break formed in another duplex. The bacterial DNA gyrase, a target for broad-spectrum antibiotics, is the sole Topo2A enzyme able to introduce negative supercoils. We reveal here for the first time the architecture of the full-length Thermus thermophilus DNA gyrase alone and in a cleavage complex with a 155 bp DNA duplex in the presence of the antibiotic ciprofloxacin, using cryo-electron microscopy. The structural organization of the subunits of the full-length DNA gyrase points to a central role of the ATPase domain acting like a 'crossover trap' that may help to sequester the DNA positive crossover before strand passage. Our structural data unveil how DNA is asymmetrically wrapped around the gyrase-specific C-terminal β-pinwheel domains and guided to introduce negative supercoils through cooperativity between the ATPase and β-pinwheel domains. The overall conformation of the drug-induced DNA binding-cleavage complex also suggests that ciprofloxacin traps a DNA pre-transport conformation. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_2361.map.gz emd_2361.map.gz | 25.1 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-2361-v30.xml emd-2361-v30.xml emd-2361.xml emd-2361.xml | 9.8 KB 9.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_2361.png emd_2361.png | 48.5 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2361 http://ftp.pdbj.org/pub/emdb/structures/EMD-2361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2361 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_2361_validation.pdf.gz emd_2361_validation.pdf.gz | 196.2 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_2361_full_validation.pdf.gz emd_2361_full_validation.pdf.gz | 195.3 KB | 表示 | |
| XML形式データ |  emd_2361_validation.xml.gz emd_2361_validation.xml.gz | 6 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2361 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2361 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2361 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2361 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_2361.map.gz / 形式: CCP4 / 大きさ: 26.4 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_2361.map.gz / 形式: CCP4 / 大きさ: 26.4 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Recontruction of the full length Thermus thermophilus DNA gyrase bound to DNA. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.92 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : DNA-bound complex of Thermus thermophilus DNA gyrase with a 155bp...
| 全体 | 名称: DNA-bound complex of Thermus thermophilus DNA gyrase with a 155bp DNA in presence of ADPNP and ciprofoxacin. |
|---|---|
| 要素 |
|
-超分子 #1000: DNA-bound complex of Thermus thermophilus DNA gyrase with a 155bp...
| 超分子 | 名称: DNA-bound complex of Thermus thermophilus DNA gyrase with a 155bp DNA in presence of ADPNP and ciprofoxacin. タイプ: sample / ID: 1000 / 詳細: monodisperse complex / 集合状態: dimer / Number unique components: 1 |
|---|---|
| 分子量 | 実験値: 413 KDa / 理論値: 413 KDa / 手法: native mass spectrometry |
-分子 #1: DNA gyrase
| 分子 | 名称: DNA gyrase / タイプ: protein_or_peptide / ID: 1 / Name.synonym: bacterial DNA topoisomerase 2A 詳細: The two subinits of the DNA gyrase were fused for structural stability. 集合状態: dimer / 組換発現: Yes |
|---|---|
| 由来(天然) | 生物種:   Thermus thermophilus (バクテリア) Thermus thermophilus (バクテリア) |
| 分子量 | 実験値: 321 KDa / 理論値: 321 KDa |
| 組換発現 | 生物種:  |
-分子 #2: linear DNA
| 分子 | 名称: linear DNA / タイプ: dna / ID: 2 詳細: A 155bp DNA was used to form the DNA-bound complex. ADPNP (a non hydrolyzable analog of ATP) and ciprofloxacin (quinolone antibiotic) were added to stabilize the complex 分類: DNA / Structure: DOUBLE HELIX / Synthetic?: No |
|---|---|
| 由来(天然) | 生物種:  |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.150 mg/mL |
|---|---|
| 緩衝液 | pH: 8 / 詳細: 20mM Hepes, 100 mM NaCl, 5mM MgCl2, 1mM DTT |
| グリッド | 詳細: Quantifoil R 2/2 holey carbon copper grids |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 95 % / チャンバー内温度: 283 K / 装置: FEI VITROBOT MARK IV / 手法: Plunging immediately after blotting |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TECNAI F30 |
|---|---|
| 日付 | 2011年12月11日 |
| 撮影 | カテゴリ: CCD / フィルム・検出器のモデル: FEI EAGLE (4k x 4k) / 実像数: 900 / 平均電子線量: 20 e/Å2 |
| 電子線 | 加速電圧: 100 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 倍率(補正後): 59000 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2 mm / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): -1.0 µm |
| 試料ステージ | 試料ホルダーモデル: GATAN LIQUID NITROGEN |
| 実験機器 |  モデル: Tecnai F30 / 画像提供: FEI Company |
- 画像解析
画像解析
| CTF補正 | 詳細: phase flipping (each particle) |
|---|---|
| 最終 再構成 | 想定した対称性 - 点群: C1 (非対称) / アルゴリズム: OTHER / 解像度のタイプ: BY AUTHOR / 解像度: 23.0 Å / 解像度の算出法: FSC 0.5 CUT-OFF / ソフトウェア - 名称: EMAN2 / 使用した粒子像数: 41500 |
 ムービー
ムービー コントローラー
コントローラー












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















