+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22131 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human CAK in complex with THZ1 | |||||||||||||||
 Map data Map data | Post-processed (sharpened, filtered) cryo-EM map, clipped to 134 pixels (as used for coordinate refinement). | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Kinase / transcription / cell cycle / complex / TRANSFERASE | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationRNA polymerase II CTD heptapeptide repeat S5 kinase activity / ventricular system development / snRNA transcription by RNA polymerase II / transcription factor TFIIK complex / CAK-ERCC2 complex / transcription factor TFIIH core complex / transcription factor TFIIH holo complex / cyclin-dependent protein serine/threonine kinase activator activity / adult heart development / [RNA-polymerase]-subunit kinase ...RNA polymerase II CTD heptapeptide repeat S5 kinase activity / ventricular system development / snRNA transcription by RNA polymerase II / transcription factor TFIIK complex / CAK-ERCC2 complex / transcription factor TFIIH core complex / transcription factor TFIIH holo complex / cyclin-dependent protein serine/threonine kinase activator activity / adult heart development / [RNA-polymerase]-subunit kinase / RNA Polymerase I Transcription Termination / cyclin-dependent protein serine/threonine kinase regulator activity / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / HIV Transcription Initiation / RNA Polymerase II HIV Promoter Escape / Transcription of the HIV genome / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA Polymerase I Transcription Initiation / RNA polymerase II transcribes snRNA genes / regulation of G1/S transition of mitotic cell cycle / cyclin-dependent kinase / cyclin-dependent protein serine/threonine kinase activity / ATP-dependent activity, acting on DNA / Tat-mediated elongation of the HIV-1 transcript / Cyclin E associated events during G1/S transition / Formation of HIV-1 elongation complex containing HIV-1 Tat / Cyclin A:Cdk2-associated events at S phase entry / Formation of HIV elongation complex in the absence of HIV Tat / cyclin-dependent protein kinase holoenzyme complex / Cyclin A/B1/B2 associated events during G2/M transition / RNA Polymerase II Transcription Elongation / Formation of RNA Pol II elongation complex / RNA polymerase II CTD heptapeptide repeat kinase activity / RNA Polymerase II Pre-transcription Events / positive regulation of smooth muscle cell proliferation / male germ cell nucleus / TP53 Regulates Transcription of DNA Repair Genes / nucleotide-excision repair / transcription initiation at RNA polymerase II promoter / RNA Polymerase I Promoter Escape / G1/S transition of mitotic cell cycle / NoRC negatively regulates rRNA expression / response to calcium ion / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / Formation of TC-NER Pre-Incision Complex / fibrillar center / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / Cyclin D associated events in G1 / kinase activity / Gap-filling DNA repair synthesis and ligation in TC-NER / RUNX1 regulates transcription of genes involved in differentiation of HSCs / transcription by RNA polymerase II / protein kinase activity / regulation of cell cycle / protein stabilization / cell division / protein serine kinase activity / DNA repair / protein serine/threonine kinase activity / regulation of transcription by RNA polymerase II / negative regulation of apoptotic process / perinuclear region of cytoplasm / positive regulation of transcription by RNA polymerase II / zinc ion binding / nucleoplasm / ATP binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||||||||
 Authors Authors | Greber BJ / Perez-Bertoldi JM | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2020 Journal: Proc Natl Acad Sci U S A / Year: 2020Title: The cryoelectron microscopy structure of the human CDK-activating kinase. Authors: Basil J Greber / Juan M Perez-Bertoldi / Kif Lim / Anthony T Iavarone / Daniel B Toso / Eva Nogales /  Abstract: The human CDK-activating kinase (CAK), a complex composed of cyclin-dependent kinase (CDK) 7, cyclin H, and MAT1, is a critical regulator of transcription initiation and the cell cycle. It acts by ...The human CDK-activating kinase (CAK), a complex composed of cyclin-dependent kinase (CDK) 7, cyclin H, and MAT1, is a critical regulator of transcription initiation and the cell cycle. It acts by phosphorylating the C-terminal heptapeptide repeat domain of the RNA polymerase II (Pol II) subunit RPB1, which is an important regulatory event in transcription initiation by Pol II, and it phosphorylates the regulatory T-loop of CDKs that control cell cycle progression. Here, we have determined the three-dimensional (3D) structure of the catalytic module of human CAK, revealing the structural basis of its assembly and providing insight into CDK7 activation in this context. The unique third component of the complex, MAT1, substantially extends the interaction interface between CDK7 and cyclin H, explaining its role as a CAK assembly factor, and it forms interactions with the CDK7 T-loop, which may contribute to enhancing CAK activity. We have also determined the structure of the CAK in complex with the covalently bound inhibitor THZ1 in order to provide insight into the binding of inhibitors at the CDK7 active site and to aid in the rational design of therapeutic compounds. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22131.map.gz emd_22131.map.gz | 8.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22131-v30.xml emd-22131-v30.xml emd-22131.xml emd-22131.xml | 22.8 KB 22.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22131.png emd_22131.png | 134.1 KB | ||
| Masks |  emd_22131_msk_1.map emd_22131_msk_1.map | 27 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-22131.cif.gz emd-22131.cif.gz | 7 KB | ||
| Others |  emd_22131_half_map_1.map.gz emd_22131_half_map_1.map.gz emd_22131_half_map_2.map.gz emd_22131_half_map_2.map.gz | 20.7 MB 20.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22131 http://ftp.pdbj.org/pub/emdb/structures/EMD-22131 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22131 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22131 | HTTPS FTP |
-Related structure data
| Related structure data |  6xd3MC  6xbzC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10438 (Title: Single-particle cryo-EM of the human CDK-activating kinase in complex with THZ1 EMPIAR-10438 (Title: Single-particle cryo-EM of the human CDK-activating kinase in complex with THZ1Data size: 2.4 TB Data #1: Unaligned movies of human CAK-THZ1 [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22131.map.gz / Format: CCP4 / Size: 9.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22131.map.gz / Format: CCP4 / Size: 9.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed (sharpened, filtered) cryo-EM map, clipped to 134 pixels (as used for coordinate refinement). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.372 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_22131_msk_1.map emd_22131_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
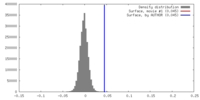
| Density Histograms |
-Half map: Unsharpened half-map (full size, 198 pixels).
| File | emd_22131_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened half-map (full size, 198 pixels). | ||||||||||||
| Projections & Slices |
| ||||||||||||
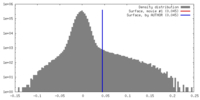
| Density Histograms |
-Half map: Unsharpened half-map (full size, 198 pixels).
| File | emd_22131_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened half-map (full size, 198 pixels). | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CDK-activating kinase assembly factor MAT1, Cyclin-H, Cyclin-depe...
| Entire | Name: CDK-activating kinase assembly factor MAT1, Cyclin-H, Cyclin-dependent kinase 7 (E.C.2.7.11.22,2.7.11.23) |
|---|---|
| Components |
|
-Supramolecule #1: CDK-activating kinase assembly factor MAT1, Cyclin-H, Cyclin-depe...
| Supramolecule | Name: CDK-activating kinase assembly factor MAT1, Cyclin-H, Cyclin-dependent kinase 7 (E.C.2.7.11.22,2.7.11.23) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: Recombinantly expressed as a trimeric complex in insect cells (MAT1 subunit truncated) and then modified with covalent inhibitor THZ1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 80 KDa |
-Macromolecule #1: CDK-activating kinase assembly factor MAT1
| Macromolecule | Name: CDK-activating kinase assembly factor MAT1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.234531 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: SNAPVTFSTG IKMGQHISLA PIHKLEEALY EYQPLQIETY GPHVPELEML GRLGYLNHVR AASPQDLAGG YTSSLACHRA LQDAFSGLF WQPS UniProtKB: CDK-activating kinase assembly factor MAT1 |
-Macromolecule #2: Cyclin-H
| Macromolecule | Name: Cyclin-H / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.695473 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MYHNSSQKRH WTFSSEEQLA RLRADANRKF RCKAVANGKV LPNDPVFLEP HEEMTLCKYY EKRLLEFCSV FKPAMPRSVV GTACMYFKR FYLNNSVMEY HPRIIMLTCA FLACKVDEFN VSSPQFVGNL RESPLGQEKA LEQILEYELL LIQQLNFHLI V HNPYRPFE ...String: MYHNSSQKRH WTFSSEEQLA RLRADANRKF RCKAVANGKV LPNDPVFLEP HEEMTLCKYY EKRLLEFCSV FKPAMPRSVV GTACMYFKR FYLNNSVMEY HPRIIMLTCA FLACKVDEFN VSSPQFVGNL RESPLGQEKA LEQILEYELL LIQQLNFHLI V HNPYRPFE GFLIDLKTRY PILENPEILR KTADDFLNRI ALTDAYLLYT PSQIALTAIL SSASRAGITM ESYLSESLML KE NRTCLSQ LLDIMKSMRN LVKKYEPPRS EEVAVLKQKL ERCHSAELAL NVITKKRKGY EDDDYVSKKS KHEEEEWTDD DLV ESL UniProtKB: Cyclin-H |
-Macromolecule #3: Cyclin-dependent kinase 7
| Macromolecule | Name: Cyclin-dependent kinase 7 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: cyclin-dependent kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.442574 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: SNAMALDVKS RAKRYEKLDF LGEGQFATVY KARDKNTNQI VAIKKIKLGH RSEAKDGINR TALREIKLLQ ELSHPNIIGL LDAFGHKSN ISLVFDFMET DLEVIIKDNS LVLTPSHIKA YMLMTLQGLE YLHQHWILHR DLKPNNLLLD ENGVLKLADF G LAKSFG(SEP)P ...String: SNAMALDVKS RAKRYEKLDF LGEGQFATVY KARDKNTNQI VAIKKIKLGH RSEAKDGINR TALREIKLLQ ELSHPNIIGL LDAFGHKSN ISLVFDFMET DLEVIIKDNS LVLTPSHIKA YMLMTLQGLE YLHQHWILHR DLKPNNLLLD ENGVLKLADF G LAKSFG(SEP)P NRAYTHQVVT RWYRAPELLF GARMYGVGVD MWAVGCILAE LLLRVPFLPG DSDLDQLTRI FETLGTPT E EQWPDMCSLP DYVTFKSFPG IPLHHIFSAA GDDLLDLIQG LFLFNPCARI TATQALKMKY FSNRPGPTPG CQLPRPNCP VETLKEQSNP ALAIKRKRTE ALEQGGLPKK LIF UniProtKB: Cyclin-dependent kinase 7 |
-Macromolecule #4: N-(3-{[5-chloro-4-(1H-indol-3-yl)pyrimidin-2-yl]amino}phenyl)-4-{...
| Macromolecule | Name: N-(3-{[5-chloro-4-(1H-indol-3-yl)pyrimidin-2-yl]amino}phenyl)-4-{[4-(dimethylamino)butanoyl]amino}benzamide type: ligand / ID: 4 / Number of copies: 1 / Formula: V0G |
|---|---|
| Molecular weight | Theoretical: 568.069 Da |
| Chemical component information | 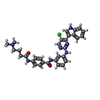 ChemComp-V0G: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.9 Component:
| ||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 5007 / Average exposure time: 2.0 sec. / Average electron dose: 69.0 e/Å2 / Details: 69 frames per movie |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 72886 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.3 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)