[English] 日本語
 Yorodumi
Yorodumi- EMDB-21306: Cryo-EM map of the C-terminal half of Leucine Rich Repeat Kinase ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21306 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of the C-terminal half of Leucine Rich Repeat Kinase 2 as a monomer at 8.1 angstroms | ||||||||||||
 Map data Map data | 8.1A map of a Leucine Rich Repeat Kinase 2 truncate as a monomer | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.1 Å | ||||||||||||
 Authors Authors | Leschziner A / Deniston C / Lahiri I | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structure of LRRK2 in Parkinson's disease and model for microtubule interaction. Authors: C K Deniston / J Salogiannis / S Mathea / D M Snead / I Lahiri / M Matyszewski / O Donosa / R Watanabe / J Böhning / A K Shiau / S Knapp / E Villa / S L Reck-Peterson / A E Leschziner /     Abstract: Leucine-rich repeat kinase 2 (LRRK2) is the most commonly mutated gene in familial Parkinson's disease and is also linked to its idiopathic form. LRRK2 has been proposed to function in membrane ...Leucine-rich repeat kinase 2 (LRRK2) is the most commonly mutated gene in familial Parkinson's disease and is also linked to its idiopathic form. LRRK2 has been proposed to function in membrane trafficking and colocalizes with microtubules. Despite the fundamental importance of LRRK2 for understanding and treating Parkinson's disease, structural information on the enzyme is limited. Here we report the structure of the catalytic half of LRRK2, and an atomic model of microtubule-associated LRRK2 built using a reported cryo-electron tomography in situ structure. We propose that the conformation of the LRRK2 kinase domain regulates its interactions with microtubules, with a closed conformation favouring oligomerization on microtubules. We show that the catalytic half of LRRK2 is sufficient for filament formation and blocks the motility of the microtubule-based motors kinesin 1 and cytoplasmic dynein 1 in vitro. Kinase inhibitors that stabilize an open conformation relieve this interference and reduce the formation of LRRK2 filaments in cells, whereas inhibitors that stabilize a closed conformation do not. Our findings suggest that LRRK2 can act as a roadblock for microtubule-based motors and have implications for the design of therapeutic LRRK2 kinase inhibitors. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21306.map.gz emd_21306.map.gz | 9.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21306-v30.xml emd-21306-v30.xml emd-21306.xml emd-21306.xml | 21.9 KB 21.9 KB | Display Display |  EMDB header EMDB header |
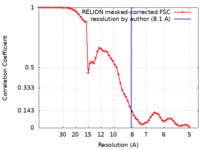
| FSC (resolution estimation) |  emd_21306_fsc.xml emd_21306_fsc.xml | 6.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_21306.png emd_21306.png | 94.2 KB | ||
| Others |  emd_21306_additional.map.gz emd_21306_additional.map.gz emd_21306_additional_1.map.gz emd_21306_additional_1.map.gz emd_21306_half_map_1.map.gz emd_21306_half_map_1.map.gz emd_21306_half_map_2.map.gz emd_21306_half_map_2.map.gz | 18.3 MB 18.3 MB 18 MB 18 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21306 http://ftp.pdbj.org/pub/emdb/structures/EMD-21306 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21306 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21306 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21306.map.gz / Format: CCP4 / Size: 19.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21306.map.gz / Format: CCP4 / Size: 19.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 8.1A map of a Leucine Rich Repeat Kinase 2 truncate as a monomer | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Sharpened 8.1A map of a Leucine Rich Repeat...
| File | emd_21306_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened 8.1A map of a Leucine Rich Repeat Kinase 2 truncate as a monomer | ||||||||||||
| Projections & Slices |
| ||||||||||||
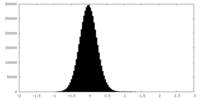
| Density Histograms |
-Additional map: Sharpened 8.1A map of a Leucine Rich Repeat...
| File | emd_21306_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened 8.1A map of a Leucine Rich Repeat Kinase 2 truncate as a monomer | ||||||||||||
| Projections & Slices |
| ||||||||||||
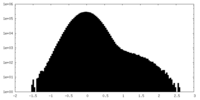
| Density Histograms |
-Half map: Half map 2
| File | emd_21306_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
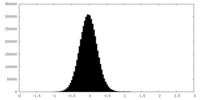
| Density Histograms |
-Half map: Half map 1
| File | emd_21306_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
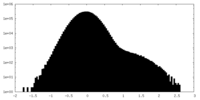
| Density Histograms |
- Sample components
Sample components
-Entire : C-terminal half of monomeric Leucine Rich Repeat Kinase 2 (LRRK2)
| Entire | Name: C-terminal half of monomeric Leucine Rich Repeat Kinase 2 (LRRK2) |
|---|---|
| Components |
|
-Supramolecule #1: C-terminal half of monomeric Leucine Rich Repeat Kinase 2 (LRRK2)
| Supramolecule | Name: C-terminal half of monomeric Leucine Rich Repeat Kinase 2 (LRRK2) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: C-terminal half consists of residue 1327 to C-terminus. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 137 KDa |
-Macromolecule #1: Leucine Rich Repeat Kinase 2
| Macromolecule | Name: Leucine Rich Repeat Kinase 2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO / EC number: non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: KKAVPYNRMK LMIVGNTGSG KTTLLQQLMK TKKSDLGMQS ATVGIDVKDW PIQIRDKRKR DLVLNVWDFA GREE FYSTH PHFMTQRALY LAVYDLSKGQ AEVDAMKPWL FNIKARASSS PVILVGTHLD VSDEKQRKAC MSKIT KELL NKRGFPAIRD YHFVNATEES ...String: KKAVPYNRMK LMIVGNTGSG KTTLLQQLMK TKKSDLGMQS ATVGIDVKDW PIQIRDKRKR DLVLNVWDFA GREE FYSTH PHFMTQRALY LAVYDLSKGQ AEVDAMKPWL FNIKARASSS PVILVGTHLD VSDEKQRKAC MSKIT KELL NKRGFPAIRD YHFVNATEES DALAKLRKTI INESLNFKIR DQLVVGQLIP DCYVELEKII LSERKN VPI EFPVIDRKRL LQLVRENQLQ LDENELPHAV HFLNESGVLL HFQDPALQLS DLYFVEPKWL CKIMAQI LT VKVEGCPKHP KGIISRRDVE KFLSKKRKFP KNYMSQYFKL LEKFQIALPI GEEYLLVPSS LSDHRPVI E LPHCENSEII IRLYEMPYFP MGFWSRLINR LLEISPYMLS GRERALRPNR MYWRQGIYLN WSPEAYCLV GSEVLDNHPE SFLKITVPSC RKGCILLGQV VDHIDSLMEE WFPGLLEIDI CGEGETLLKK WALYSFNDGE EHQKILLDD LMKKAEEGDL LVNPDQPRLT IPISQIAPDL ILADLPRNIM LNNDELEFEQ APEFLLGDGS F GSVYRAAY EGEEVAVKIF NKHTSLRLLR QELVVLCHLH HPSLISLLAA GIRPRMLVME LASKGSLDRL LQ QDKASLT RTLQHRIALH VADGLRYLHS AMIIYRDLKP HNVLLFTLYP NAAIIAKIAD YGIAQYCCRM GIK TSEGTP GFRAPEVARG NVIYNQQADV YSFGLLLYDI LTTGGRIVEG LKFPNEFDEL EIQGKLPDPV KEYG CAPWP MVEKLIKQCL KENPQERPTS AQVFDILNSA ELVCLTRRIL LPKNVIVECM VATHHNSRNA SIWLG CGHT DRGQLSFLDL NTEGYTSEEV ADSRILCLAL VHLPVEKESW IVSGTQSGTL LVINTEDGKK RHTLEK MTD SVTCLYCNSF SKQSKQKNFL LVGTADGKLA IFEDKTVKLK GAAPLKILNI GNVSTPLMCL SESTNST ER NVMWGGCGTK IFSFSNDFTI QKLIETRTSQ LFSYAAFSDS NIITVVVDTA LYIAKQNSPV VEVWDKKT E KLCGLIDCVH FLREVMVKEN KESKHKMSYS GRVKTLCLQK NTALWIGTGG GHILLLDLST RRLIRVIYN FCNSVRVMMT AQLGSLKNVM LVLGYNRKNT EGTQKQKEIQ SCLTVWDINL PHEVQNLEKH IEVRKELAEK MRRTSVE |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||||||||
| Grid | Model: Quantifoil, UltrAuFoil, R1.2/1.3 / Material: COPPER/RHODIUM / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK II | |||||||||||||||||||||
| Details | This sample was imaged over a number of datasets. Specimen concentrations ranged from 0.14(1uM)-0.84(6uM) mg/mL, however the average was 0.5 and thus was entered as the reported specimen concentration. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Details | Datasets imaged at 36kx, either counting or super resolution mode) magnifications were collected. One dataset collected at 30 degree tilt. |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 7067 / Average electron dose: 4.2 e/Å2 Details: Datasets collected at 36kx, either counting or super resolution mod,) magnifications were collected. Only a single dose has been entered, however datasets were collected at various doses ...Details: Datasets collected at 36kx, either counting or super resolution mod,) magnifications were collected. Only a single dose has been entered, however datasets were collected at various doses between 4.2 and 10 electrons per angstrom squared. See method section of paper for more details. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)