[English] 日本語
 Yorodumi
Yorodumi- EMDB-21221: CryoEM map of Hrd1-Usa1/Der1/Hrd3 complex of the expected topology -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21221 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM map of Hrd1-Usa1/Der1/Hrd3 complex of the expected topology | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | retro-translocation / ERAD / protein degradation / ubiquitination / PROTEIN TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationHrd1p ubiquitin ligase ERAD-M complex / detection of unfolded protein / luminal surveillance complex / Hrd1p ubiquitin ligase complex / misfolded protein transport / Hrd1p ubiquitin ligase ERAD-L complex / fungal-type cell wall organization / signal recognition particle binding / negative regulation of protein autoubiquitination / misfolded protein binding ...Hrd1p ubiquitin ligase ERAD-M complex / detection of unfolded protein / luminal surveillance complex / Hrd1p ubiquitin ligase complex / misfolded protein transport / Hrd1p ubiquitin ligase ERAD-L complex / fungal-type cell wall organization / signal recognition particle binding / negative regulation of protein autoubiquitination / misfolded protein binding / positive regulation of protein autoubiquitination / retrograde protein transport, ER to cytosol / protein quality control for misfolded or incompletely synthesized proteins / protein K48-linked ubiquitination / endoplasmic reticulum unfolded protein response / protein autoubiquitination / ERAD pathway / mRNA splicing, via spliceosome / RING-type E3 ubiquitin transferase / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / ubiquitin-dependent protein catabolic process / molecular adaptor activity / endoplasmic reticulum membrane / endoplasmic reticulum / zinc ion binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | |||||||||
 Authors Authors | Wu X / Rapoport TA | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2020 Journal: Science / Year: 2020Title: Structural basis of ER-associated protein degradation mediated by the Hrd1 ubiquitin ligase complex. Authors: Xudong Wu / Marc Siggel / Sergey Ovchinnikov / Wei Mi / Vladimir Svetlov / Evgeny Nudler / Maofu Liao / Gerhard Hummer / Tom A Rapoport /   Abstract: Misfolded luminal endoplasmic reticulum (ER) proteins undergo ER-associated degradation (ERAD-L): They are retrotranslocated into the cytosol, polyubiquitinated, and degraded by the proteasome. ERAD- ...Misfolded luminal endoplasmic reticulum (ER) proteins undergo ER-associated degradation (ERAD-L): They are retrotranslocated into the cytosol, polyubiquitinated, and degraded by the proteasome. ERAD-L is mediated by the Hrd1 complex (composed of Hrd1, Hrd3, Der1, Usa1, and Yos9), but the mechanism of retrotranslocation remains mysterious. Here, we report a structure of the active Hrd1 complex, as determined by cryo-electron microscopy analysis of two subcomplexes. Hrd3 and Yos9 jointly create a luminal binding site that recognizes glycosylated substrates. Hrd1 and the rhomboid-like Der1 protein form two "half-channels" with cytosolic and luminal cavities, respectively, and lateral gates facing one another in a thinned membrane region. These structures, along with crosslinking and molecular dynamics simulation results, suggest how a polypeptide loop of an ERAD-L substrate moves through the ER membrane. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21221.map.gz emd_21221.map.gz | 14.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21221-v30.xml emd-21221-v30.xml emd-21221.xml emd-21221.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21221.png emd_21221.png | 102.1 KB | ||
| Filedesc metadata |  emd-21221.cif.gz emd-21221.cif.gz | 6.6 KB | ||
| Others |  emd_21221_additional.map.gz emd_21221_additional.map.gz | 11.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21221 http://ftp.pdbj.org/pub/emdb/structures/EMD-21221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21221 | HTTPS FTP |
-Validation report
| Summary document |  emd_21221_validation.pdf.gz emd_21221_validation.pdf.gz | 533.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21221_full_validation.pdf.gz emd_21221_full_validation.pdf.gz | 533.3 KB | Display | |
| Data in XML |  emd_21221_validation.xml.gz emd_21221_validation.xml.gz | 5.3 KB | Display | |
| Data in CIF |  emd_21221_validation.cif.gz emd_21221_validation.cif.gz | 6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21221 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21221 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21221 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21221 | HTTPS FTP |
-Related structure data
| Related structure data |  6vjzMC  6vjyC  6vk0C  6vk1C  6vk3C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21221.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21221.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unsharpened map
| File | emd_21221_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
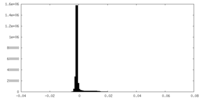
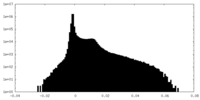
| Density Histograms |
- Sample components
Sample components
-Entire : complex of Hrd1-Usa1/Der1/Hrd3 in the expected topology
| Entire | Name: complex of Hrd1-Usa1/Der1/Hrd3 in the expected topology |
|---|---|
| Components |
|
-Supramolecule #1: complex of Hrd1-Usa1/Der1/Hrd3 in the expected topology
| Supramolecule | Name: complex of Hrd1-Usa1/Der1/Hrd3 in the expected topology type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Degradation in the endoplasmic reticulum protein 1
| Macromolecule | Name: Degradation in the endoplasmic reticulum protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.411715 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDAVILNLLG DIPLVTRLWT IGCLVLSGLT SLRIVDPGKV VYSYDLVFKK GQYGRLLYSI FDYGAFNWIS MINIFVSANH LSTLENSFN LRRKFCWIIF LLLVILVKMT SIEQPAASLG VLLHENLVYY ELKKNGNQMN VRFFGAIDVS PSIFPIYMNA V MYFVYKRS ...String: MDAVILNLLG DIPLVTRLWT IGCLVLSGLT SLRIVDPGKV VYSYDLVFKK GQYGRLLYSI FDYGAFNWIS MINIFVSANH LSTLENSFN LRRKFCWIIF LLLVILVKMT SIEQPAASLG VLLHENLVYY ELKKNGNQMN VRFFGAIDVS PSIFPIYMNA V MYFVYKRS WLEIAMNFMP GHVIYYMDDI IGKIYGIDLC KSPYDWFRNT ETP UniProtKB: Degradation in the endoplasmic reticulum protein 1 |
-Macromolecule #2: ERAD-associated E3 ubiquitin-protein ligase HRD1
| Macromolecule | Name: ERAD-associated E3 ubiquitin-protein ligase HRD1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: RING-type E3 ubiquitin transferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 55.638773 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVPENRRKQL AIFVVVTYLL TFYCVYSATK TSVSFLQVTL KLNEGFNLMV LSIFILLNST LLWQLLTKLL FGELRLIEHE HIFERLPFT IINTLFMSSL FHERYFFTVA FFGLLLLYLK VFHWILKDRL EALLQSINDS TTMKTLIFSR FSFNLVLLAV V DYQIITRC ...String: MVPENRRKQL AIFVVVTYLL TFYCVYSATK TSVSFLQVTL KLNEGFNLMV LSIFILLNST LLWQLLTKLL FGELRLIEHE HIFERLPFT IINTLFMSSL FHERYFFTVA FFGLLLLYLK VFHWILKDRL EALLQSINDS TTMKTLIFSR FSFNLVLLAV V DYQIITRC ISSIYTNQKS DIESTSLYLI QVMEFTMLLI DLLNLFLQTC LNFWEFYRSQ QSLSNENNHI VHGDPTDENT VE SDQSQPV LNDDDDDDDD DRQFTGLEGK FMYEKAIDVF TRFLKTALHL SMLIPFRMPM MLLKDVVWDI LALYQSGTSL WKI WRNNKQ LDDTLVTVTV EQLQNSANDD NICIICMDEL IHSPNQQTWK NKNKKPKRLP CGHILHLSCL KNWMERSQTC PICR LPVFD EKGNVVQTTF TSNSDITTQT TVTDSTGIAT DQQGFANEVD LLPTRTTSPD IRIVPTQNID TLAMRTRSTS TPSPT UniProtKB: ERAD-associated E3 ubiquitin-protein ligase HRD1 |
-Macromolecule #3: ERAD-associated E3 ubiquitin-protein ligase component HRD3
| Macromolecule | Name: ERAD-associated E3 ubiquitin-protein ligase component HRD3 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 88.239102 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MITLLLYLCV ICNAIVLIRA DSIADPWPEA RHLLNTIAKS RDPMKEAAME PNADEFVGFY VPMDYSPRNE EKNYQSIWQN EITDSQRHI YELLVQSSEQ FNNSEATYTL SQIHLWSQYN FPHNMTLAHK YLEKFNDLTH FTNHSAIFDL AVMYATGGCA S GNDQTVIP ...String: MITLLLYLCV ICNAIVLIRA DSIADPWPEA RHLLNTIAKS RDPMKEAAME PNADEFVGFY VPMDYSPRNE EKNYQSIWQN EITDSQRHI YELLVQSSEQ FNNSEATYTL SQIHLWSQYN FPHNMTLAHK YLEKFNDLTH FTNHSAIFDL AVMYATGGCA S GNDQTVIP QDSAKALLYY QRAAQLGNLK AKQVLAYKYY SGFNVPRNFH KSLVLYRDIA EQLRKSYSRD EWDIVFPYWE SY NVRISDF ESGLLGKGLN SVPSSTVRKR TTRPDIGSPF IAQVNGVQMT LQIEPMGRFA FNGNDGNING DEDDEDASER RII RIYYAA LNDYKGTYSQ SRNCERAKNL LELTYKEFQP HVDNLDPLQV FYYVRCLQLL GHMYFTGEGS SKPNIHMAEE ILTT SLEIS RRAQGPIGRA CIDLGLINQY ITNNISQAIS YYMKAMKTQA NNGIVEFQLS KLATSFPEEK IGDPFNLMET AYLNG FIPA IYEFAVMIES GMNSKSSVEN TAYLFKTFVD KNEAIMAPKL RTAFAALIND RSEVALWAYS QLAEQGYETA QVSAAY LMY QLPYEFEDPP RTTDQRKTLA ISYYTRAFKQ GNIDAGVVAG DIYFQMQNYS KAMALYQGAA LKYSIQAIWN LGYMHEH GL GVNRDFHLAK RYYDQVSEHD HRFYLASKLS VLKLHLKSWL TWITREKVNY WKPSSPLNPN EDTQHSKTSW YKQLTKIL Q RMRHKEDSDK AAEDSHKHRT VVQNGANHRG DDQEEASEIL GFQMED UniProtKB: ERAD-associated E3 ubiquitin-protein ligase component HRD3 |
-Macromolecule #4: U1 SNP1-associating protein 1
| Macromolecule | Name: U1 SNP1-associating protein 1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 39.944301 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VRAADNTSSA NDNNTVENDE SAWNRRVVRP LRNSFPLLLV LIRTFYLIGY NSLVPFFIIL EFGSFLPWKY IILLSLLFIF RTVWNTQEV WNLWRDYLHL NEIDEVKFSQ IKEFINSNSL TLNFYKKCKD TQSAIDLLMI PNLHEQRLSV YSKYDIEYDT N TPDVGQLN ...String: VRAADNTSSA NDNNTVENDE SAWNRRVVRP LRNSFPLLLV LIRTFYLIGY NSLVPFFIIL EFGSFLPWKY IILLSLLFIF RTVWNTQEV WNLWRDYLHL NEIDEVKFSQ IKEFINSNSL TLNFYKKCKD TQSAIDLLMI PNLHEQRLSV YSKYDIEYDT N TPDVGQLN LLFIKVLSGE IPKDALDELF KEFFELYETT RNMNTLYPQD SLNELLLMIW KESQKKDINT LPKYRRWFQT LC SQIAEHN VLDVVLRYII PDPVNDRVIT AVIKNFVLFW VTLLPYVKEK LDDIVAQRAR DREQPAPSAQ QQENEDEALI IPD EEEPTA TGAQPHLYIP DED UniProtKB: U1 SNP1-associating protein 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 54.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: RANDOM CONICAL TILT |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.3 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.0) / Number images used: 172291 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.0) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.0) |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)