+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20525 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | ChsH1-ChsH2-Ltp2 complex | |||||||||
 Map data Map data | ChsH1-ChsH2-Ltp2 complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | an enoyl-CoA hydratase retro-aldolase complex / cholesterol metabolism / STRUCTURAL PROTEIN | |||||||||
| Biological species |  | |||||||||
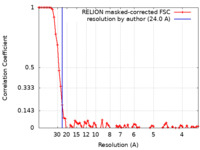
| Method | single particle reconstruction / negative staining / Resolution: 24.0 Å | |||||||||
 Authors Authors | Yang M / Yuan T / Gehring K / Sampson N | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Biochemistry / Year: 2019 Journal: Biochemistry / Year: 2019Title: Exploits a Heterohexameric Enoyl-CoA Hydratase Retro-Aldolase Complex for Cholesterol Catabolism. Authors: Tianao Yuan / Meng Yang / Kalle Gehring / Nicole S Sampson /   Abstract: Cholesterol catabolism plays an important role in 's ('s) survival and persistence in the host. exploits three β-oxidation cycles to fully degrade the side chain of cholesterol. Five cistronic ...Cholesterol catabolism plays an important role in 's ('s) survival and persistence in the host. exploits three β-oxidation cycles to fully degrade the side chain of cholesterol. Five cistronic genes in a single operon encode three enzymes, 3-oxo-4-pregnene-20-carboxyl-CoA dehydrogenase (ChsE1-ChsE2), 3-oxo-4,17-pregnadiene-20-carboxyl-CoA hydratase (ChsH1-ChsH2), and 17-hydroxy-3-oxo-4-pregnene-20-carboxyl-CoA retro-aldolase (Ltp2), to perform the last β-oxidation cycle in this pathway. Among these three enzymes, ChsH1-ChsH2 and Ltp2 form a protein complex that is required for the catalysis of carbon-carbon bond cleavage. In this work, we report the structure of the full length ChsH1-ChsH2-Ltp2 complex based on small-angle X-ray scattering and single-particle electron microscopy data. Mutagenesis experiments confirm the requirement for Ltp2 to catalyze the retro-aldol reaction. The structure illustrates how acyl transfer between enzymes may occur. Each protomer of the ChsH1-ChsH2-Ltp2 complex contains three protein components: a chain of ChsH1, a chain of ChsH2, and a chain of Ltp2. Two protomers dimerize at the interface of Ltp2 to form a heterohexameric structure. This unique heterohexameric structure of the ChsH1-ChsH2-Ltp2 complex provides entry to further understand the mechanism of cholesterol catabolism in . | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20525.map.gz emd_20525.map.gz | 22.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20525-v30.xml emd-20525-v30.xml emd-20525.xml emd-20525.xml | 11.8 KB 11.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20525_fsc.xml emd_20525_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_20525.png emd_20525.png | 16.3 KB | ||
| Others |  emd_20525_half_map_1.map.gz emd_20525_half_map_1.map.gz emd_20525_half_map_2.map.gz emd_20525_half_map_2.map.gz | 22.4 MB 22.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20525 http://ftp.pdbj.org/pub/emdb/structures/EMD-20525 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20525 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20525 | HTTPS FTP |
-Validation report
| Summary document |  emd_20525_validation.pdf.gz emd_20525_validation.pdf.gz | 553.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20525_full_validation.pdf.gz emd_20525_full_validation.pdf.gz | 552.9 KB | Display | |
| Data in XML |  emd_20525_validation.xml.gz emd_20525_validation.xml.gz | 12.8 KB | Display | |
| Data in CIF |  emd_20525_validation.cif.gz emd_20525_validation.cif.gz | 17.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20525 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20525 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20525 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20525 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20525.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20525.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ChsH1-ChsH2-Ltp2 complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.8 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: half map 2
| File | emd_20525_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
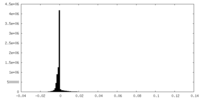
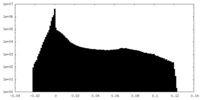
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_20525_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
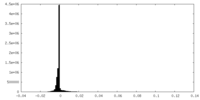
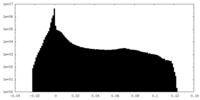
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : an enoyl-CoA hydratase retro-aldolase complex
| Entire | Name: an enoyl-CoA hydratase retro-aldolase complex |
|---|---|
| Components |
|
-Supramolecule #1: an enoyl-CoA hydratase retro-aldolase complex
| Supramolecule | Name: an enoyl-CoA hydratase retro-aldolase complex / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 180 KDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Staining | Type: NEGATIVE / Material: Uranyl Acetate |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: GATAN MULTISCAN / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)