+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20254 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Drosophila P element transposase strand transfer complex | |||||||||
 Map data Map data | symmetric (C2) cryo-EM reconstruction of the P element transposase strand-transfer complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transposase / Strand Transfer Complex / TRANSFERASE-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationP-element binding / transposase activity / DNA transposition / DNA integration / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / transferase activity / zinc ion binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Kellogg EH / Nogales E | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2019 Journal: Nat Struct Mol Biol / Year: 2019Title: Structure of a P element transposase-DNA complex reveals unusual DNA structures and GTP-DNA contacts. Authors: George E Ghanim / Elizabeth H Kellogg / Eva Nogales / Donald C Rio /  Abstract: P element transposase catalyzes the mobility of P element DNA transposons within the Drosophila genome. P element transposase exhibits several unique properties, including the requirement for a ...P element transposase catalyzes the mobility of P element DNA transposons within the Drosophila genome. P element transposase exhibits several unique properties, including the requirement for a guanosine triphosphate cofactor and the generation of long staggered DNA breaks during transposition. To gain insights into these features, we determined the atomic structure of the Drosophila P element transposase strand transfer complex using cryo-EM. The structure of this post-transposition nucleoprotein complex reveals that the terminal single-stranded transposon DNA adopts unusual A-form and distorted B-form helical geometries that are stabilized by extensive protein-DNA interactions. Additionally, we infer that the bound guanosine triphosphate cofactor interacts with the terminal base of the transposon DNA, apparently to position the P element DNA for catalysis. Our structure provides the first view of the P element transposase superfamily, offers new insights into P element transposition and implies a transposition pathway fundamentally distinct from other cut-and-paste DNA transposases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20254.map.gz emd_20254.map.gz | 4.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20254-v30.xml emd-20254-v30.xml emd-20254.xml emd-20254.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20254.png emd_20254.png | 122.6 KB | ||
| Filedesc metadata |  emd-20254.cif.gz emd-20254.cif.gz | 7 KB | ||
| Others |  emd_20254_half_map_1.map.gz emd_20254_half_map_1.map.gz emd_20254_half_map_2.map.gz emd_20254_half_map_2.map.gz | 48.4 MB 48.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20254 http://ftp.pdbj.org/pub/emdb/structures/EMD-20254 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20254 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20254 | HTTPS FTP |
-Validation report
| Summary document |  emd_20254_validation.pdf.gz emd_20254_validation.pdf.gz | 648.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20254_full_validation.pdf.gz emd_20254_full_validation.pdf.gz | 647.8 KB | Display | |
| Data in XML |  emd_20254_validation.xml.gz emd_20254_validation.xml.gz | 12.1 KB | Display | |
| Data in CIF |  emd_20254_validation.cif.gz emd_20254_validation.cif.gz | 14.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20254 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20254 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20254 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20254 | HTTPS FTP |
-Related structure data
| Related structure data |  6p5aMC  6pe2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20254.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20254.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | symmetric (C2) cryo-EM reconstruction of the P element transposase strand-transfer complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
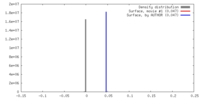
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: half map (1) of final cryo-EM reconstruction of...
| File | emd_20254_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map (1) of final cryo-EM reconstruction of the P element transposase strand-transfer complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
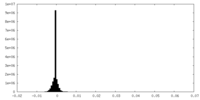
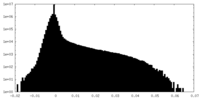
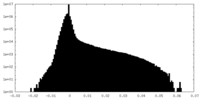
| Density Histograms |
-Half map: half map (2) of final cryo-EM reconstruction of...
| File | emd_20254_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map (2) of final cryo-EM reconstruction of the P element transposase strand-transfer complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
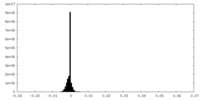
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of P element transposase in complex with strand t...
| Entire | Name: Ternary complex of P element transposase in complex with strand transfer DNA |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of P element transposase in complex with strand t...
| Supramolecule | Name: Ternary complex of P element transposase in complex with strand transfer DNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transposable element P transposase
| Macromolecule | Name: Transposable element P transposase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO EC number: Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 65.525992 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKYCKFCCKA VTGVKLIHVP KCAIKRKLWE QSLGCSLGEN SQICDTHFND SQWKAAPAKG QTFKRRRLNA DAVPSKVIEP EPEKIKEGY TSGSTQTESC SLFNENKSLR EKIRTLEYEM RRLEQQLRES QQLEESLRKI FTDTQIRILK NGGQRATFNS D DISTAICL ...String: MKYCKFCCKA VTGVKLIHVP KCAIKRKLWE QSLGCSLGEN SQICDTHFND SQWKAAPAKG QTFKRRRLNA DAVPSKVIEP EPEKIKEGY TSGSTQTESC SLFNENKSLR EKIRTLEYEM RRLEQQLRES QQLEESLRKI FTDTQIRILK NGGQRATFNS D DISTAICL HTAGPRAYNH LYKKGFPLPS RTTLYRWLSD VDIKRGCLDV VIDLMDSDGV DDADKLCVLA FDEMKVAAAF EY DSSADIV YEPSDYVQLA IVRGLKKSWK QPVFFDFNTR MDPDTLNNIL RKLHRKGYLV VAIVSDLGTG NQKLWTELGI SES KTWFSH PADDHLKIFV FSDTPHLIKL VRNHYVDSGL TINGKKLTKK TIQEALHLCN KSDLSILFKI NENHINVRSL AKQK VKLAT QLFSNTTASS IRRCYSLGYD IENATETADF FKLMNDWFDI FNSKLSTSNC IECSQPYGKQ LDIQNDILNR MSEIM RTGI LDKPKRLPFQ KGIIVNNASL DGLYKYLQEN FSMQYILTSR LNQDIVEHFF GSMRSRGGQF DHPTPLQFKY RLRKYI IAR NTEMLR UniProtKB: Transposable element P transposase |
-Macromolecule #2: Transposable element P transposase
| Macromolecule | Name: Transposable element P transposase / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO EC number: Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.086797 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: TEMDELTEDA MEYIAGYVIK KLRISDKVKE NLTFTYVDEV SHGGLIKPSE KFQEKLKELE CIFLHYTNNN NFEITNNVKE KLILAARNV DVDKQVKSFY FKIRIYFRIK YFNKKIEIKN QKQKLIGNSK LLKIKL UniProtKB: Transposable element P transposase |
-Macromolecule #3: DNA (5'-D(P*CP*GP*AP*AP*CP*TP*AP*TP*A)-3')
| Macromolecule | Name: DNA (5'-D(P*CP*GP*AP*AP*CP*TP*AP*TP*A)-3') / type: dna / ID: 3 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.827157 KDa |
| Sequence | String: (DC)(DC)(DG)(DC)(DT)(DC)(DA)(DC)(DG)(DA) (DA)(DC)(DT)(DA)(DT)(DA) |
-Macromolecule #4: DNA (5'-D(P*AP*GP*GP*TP*GP*GP*TP*CP*CP*CP*GP*TP*CP*GP*G)-3')
| Macromolecule | Name: DNA (5'-D(P*AP*GP*GP*TP*GP*GP*TP*CP*CP*CP*GP*TP*CP*GP*G)-3') type: dna / ID: 4 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.704529 KDa |
| Sequence | String: (DA)(DG)(DG)(DT)(DG)(DG)(DT)(DC)(DC)(DC) (DG)(DT)(DC)(DG)(DG)(DC)(DA)(DA)(DG)(DA) (DG)(DA)(DC)(DA)(DT)(DC)(DC)(DA)(DC) (DT)(DT)(DA)(DA)(DC)(DG)(DT)(DA)(DT) |
-Macromolecule #5: DNA (44-MER)
| Macromolecule | Name: DNA (44-MER) / type: dna / ID: 5 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.400576 KDa |
| Sequence | String: (DA)(DT)(DA)(DC)(DG)(DT)(DT)(DA)(DA)(DG) (DT)(DG)(DG)(DA)(DT)(DG)(DT)(DC)(DT)(DC) (DT)(DT)(DG)(DC)(DC)(DG)(DA)(DC)(DG) (DG)(DG)(DA)(DC)(DC)(DA)(DC)(DC)(DT)(DT) (DA) (DT)(DG)(DT)(DT)(DA)(DT) ...String: (DA)(DT)(DA)(DC)(DG)(DT)(DT)(DA)(DA)(DG) (DT)(DG)(DG)(DA)(DT)(DG)(DT)(DC)(DT)(DC) (DT)(DT)(DG)(DC)(DC)(DG)(DA)(DC)(DG) (DG)(DG)(DA)(DC)(DC)(DA)(DC)(DC)(DT)(DT) (DA) (DT)(DG)(DT)(DT)(DA)(DT)(DT)(DT) (DC)(DA)(DT)(DC)(DA)(DT)(DG)(DG)(DT)(DC) (DC)(DG) (DG)(DA)(DC)(DT)(DA)(DT)(DA) (DG)(DT)(DT)(DC)(DG)(DT)(DG)(DA)(DG)(DC) (DG)(DG) |
-Macromolecule #6: GUANOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 2 / Formula: GTP |
|---|---|
| Molecular weight | Theoretical: 523.18 Da |
| Chemical component information |  ChemComp-GTP: |
-Macromolecule #7: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 7 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Component:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: UltrAuFoil / Material: GOLD / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: 4 microliters of concentrated STC complex was applied to a Quantifoil 1.2/1.3 UltraAuFoil grid. After a 30 second incubation, the sample was blotted using blot force of 8 pN and a blot time of 6 seconds.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Details | Dataset collected at 40 degree tilt. |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 7420 pixel / Digitization - Dimensions - Height: 7676 pixel / Number grids imaged: 1 / Number real images: 1857 / Average exposure time: 10.0 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE / Details: cryosparc |
|---|---|
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: C2 (2 fold cyclic) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.0) / Software - details: beta / Number images used: 253209 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.0) / Software - details: beta version |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 100 / Target criteria: correlation coefficient |
|---|---|
| Output model |  PDB-6p5a: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)