[English] 日本語
 Yorodumi
Yorodumi- EMDB-1964: Structural and Functional Studies of LRP6 Ectodomain Reveal a Pla... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1964 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structural and Functional Studies of LRP6 Ectodomain Reveal a Platform for Wnt Signaling | |||||||||
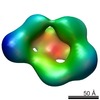
 Map data Map data | This is a surface rendering of the LRP6 ectodomain filtered to 25 angstroms resolution. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | LDL-receptor-related protein 6 / Wnt signaling pathway / Wnt co-receptor | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 25.0 Å | |||||||||
 Authors Authors | Chen S / Bubeck D / MacDonald BT / Liang WX / Mao JH / Malinauskas T / Llorca O / Aricescu AR / Siebold C / He X / Jones EY | |||||||||
 Citation Citation |  Journal: Dev Cell / Year: 2011 Journal: Dev Cell / Year: 2011Title: Structural and functional studies of LRP6 ectodomain reveal a platform for Wnt signaling. Authors: Shuo Chen / Doryen Bubeck / Bryan T MacDonald / Wen-Xue Liang / Jian-Hua Mao / Tomas Malinauskas / Oscar Llorca / A Radu Aricescu / Christian Siebold / Xi He / E Yvonne Jones /  Abstract: LDL-receptor-related protein 6 (LRP6), alongside Frizzled receptors, transduces Wnt signaling across the plasma membrane. The LRP6 ectodomain comprises four tandem β-propeller-EGF-like domain (PE) ...LDL-receptor-related protein 6 (LRP6), alongside Frizzled receptors, transduces Wnt signaling across the plasma membrane. The LRP6 ectodomain comprises four tandem β-propeller-EGF-like domain (PE) pairs that harbor binding sites for Wnt morphogens and their antagonists including Dickkopf 1 (Dkk1). To understand how these multiple interactions are integrated, we combined crystallographic analysis of the third and fourth PE pairs with electron microscopy (EM) to determine the complete ectodomain structure. An extensive inter-pair interface, conserved for the first-to-second and third-to-fourth PE interactions, contributes to a compact platform-like architecture, which is disrupted by mutations implicated in developmental diseases. EM reconstruction of the LRP6 platform bound to chaperone Mesd exemplifies a binding mode spanning PE pairs. Cellular and binding assays identify overlapping Wnt3a- and Dkk1-binding surfaces on the third PE pair, consistent with steric competition, but also suggest a model in which the platform structure supports an interplay of ligands through multiple interaction sites. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1964.map.gz emd_1964.map.gz | 791.8 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1964-v30.xml emd-1964-v30.xml emd-1964.xml emd-1964.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  1964_LRP6_2.jpg 1964_LRP6_2.jpg | 33.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1964 http://ftp.pdbj.org/pub/emdb/structures/EMD-1964 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1964 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1964 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1964.map.gz / Format: CCP4 / Size: 825.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1964.map.gz / Format: CCP4 / Size: 825.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is a surface rendering of the LRP6 ectodomain filtered to 25 angstroms resolution. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.56 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ectodomain of LRP6 residues 20 to 1361
| Entire | Name: Ectodomain of LRP6 residues 20 to 1361 |
|---|---|
| Components |
|
-Supramolecule #1000: Ectodomain of LRP6 residues 20 to 1361
| Supramolecule | Name: Ectodomain of LRP6 residues 20 to 1361 / type: sample / ID: 1000 / Details: The sample was monodisperse / Oligomeric state: Monomer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 165 KDa / Method: Multi angle light scattering |
-Macromolecule #1: LDL-receptor-related protein 6
| Macromolecule | Name: LDL-receptor-related protein 6 / type: protein_or_peptide / ID: 1 / Name.synonym: LRP6 / Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Experimental: 165 KDa |
| Recombinant expression | Organism: Homo sapiens embryonic kidney cells / Recombinant plasmid: pHLsec vector |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.030 mg/mL |
|---|---|
| Buffer | pH: 5 / Details: 150 mM NaCl, 50 mM NaAc |
| Staining | Type: NEGATIVE Details: Grids were negatively stained with 0.75% uranyl formate using the two-drop method |
| Grid | Details: carbon-coated copper grids |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 1230 |
|---|---|
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Digitization - Sampling interval: 4.56 µm / Average electron dose: 10 e/Å2 |
| Electron beam | Acceleration voltage: 100 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.9 mm / Nominal magnification: 72500 |
| Sample stage | Specimen holder: single-tilt / Specimen holder model: JEOL |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 25.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN, XMIPP / Number images used: 6999 |
|---|
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Coot Coot |
| Details | Protocol: Rigid Body. A homology model of LRP6 residues 20 to 1244 based on the LRP6P3E3P4E4 crystal structure was built using Modeller. This single rigid-body was manually docked into the electron microscopy reconstruction and subjected to automated real-space refinement in Coot. Residues linking E2 and P3 residues 628 to 632 were removed. The model was subsequently refined as two rigid bodies residues 20 to 627 and residues 633 to 1244 using Coot. Refined models were scored against the electron microscopy map using a real space correlation coefficient computing using the Bsoft package . |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















