[English] 日本語
 Yorodumi
Yorodumi- EMDB-17592: Sub-tomogram average of the open conformation of the Nap adhesion... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Sub-tomogram average of the open conformation of the Nap adhesion complex from the human pathogen Mycoplasma genitalium. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Adhesion / Mycoplasma genitalium / CELL ADHESION | |||||||||
| Function / homology |  Function and homology information Function and homology informationadhesion of symbiont to microvasculature / cell adhesion / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Mycoplasmoides genitalium G37 (bacteria) Mycoplasmoides genitalium G37 (bacteria) | |||||||||
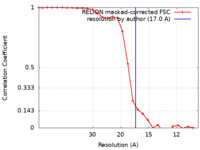
| Method | subtomogram averaging / cryo EM / Resolution: 17.0 Å | |||||||||
 Authors Authors | Sprankel L / Scheffer MP / Frangakis AF | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2023 Journal: PLoS Pathog / Year: 2023Title: Cryo-electron tomography reveals the binding and release states of the major adhesion complex from Mycoplasma genitalium. Authors: Lasse Sprankel / Margot P Scheffer / Sina Manger / Utz H Ermel / Achilleas S Frangakis /  Abstract: The nap particle is an immunogenic surface adhesion complex from Mycoplasma genitalium. It is essential for motility and responsible for binding sialylated oligosaccharides on the surface of the host ...The nap particle is an immunogenic surface adhesion complex from Mycoplasma genitalium. It is essential for motility and responsible for binding sialylated oligosaccharides on the surface of the host cell. The nap particle is composed of two P140-P110 heterodimers, the structure of which was recently solved. However, the interpretation of the mechanism by which the mycoplasma cells orchestrate adhesion remained challenging. Here, we provide cryo-electron tomography structures at ~11 Å resolution, which allow for the distinction between the bound and released state of the nap particle, displaying the in vivo conformational states. Fitting of the atomically resolved structures reveals that bound sialylated oligosaccharides are stabilized by both P110 and P140. Movement of the stalk domains allows for the transfer of conformational changes from the interior of the cell to the binding pocket, thus having the capability of an active release process. It is likely that the same mechanism can be transferred to other Mycoplasma species that belong to the pneumoniae cluster. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17592.map.gz emd_17592.map.gz | 940.1 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17592-v30.xml emd-17592-v30.xml emd-17592.xml emd-17592.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17592_fsc.xml emd_17592_fsc.xml | 2.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_17592.png emd_17592.png | 176.7 KB | ||
| Masks |  emd_17592_msk_1.map emd_17592_msk_1.map | 1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17592.cif.gz emd-17592.cif.gz | 7.1 KB | ||
| Others |  emd_17592_half_map_1.map.gz emd_17592_half_map_1.map.gz emd_17592_half_map_2.map.gz emd_17592_half_map_2.map.gz | 942.1 KB 941.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17592 http://ftp.pdbj.org/pub/emdb/structures/EMD-17592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17592 | HTTPS FTP |
-Related structure data
| Related structure data |  8pc0MC  8pbxC  8pbyC  8pbzC  8pc1C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17592.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17592.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.2 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17592_msk_1.map emd_17592_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
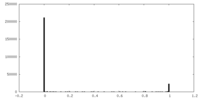
| Density Histograms |
-Half map: #1
| File | emd_17592_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
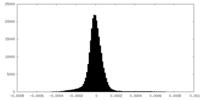
| Density Histograms |
-Half map: #2
| File | emd_17592_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
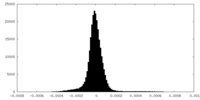
| Density Histograms |
- Sample components
Sample components
-Entire : Mildly lysed M. genitalium cells
| Entire | Name: Mildly lysed M. genitalium cells |
|---|---|
| Components |
|
-Supramolecule #1: Mildly lysed M. genitalium cells
| Supramolecule | Name: Mildly lysed M. genitalium cells / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Mycoplasmoides genitalium G37 (bacteria) Mycoplasmoides genitalium G37 (bacteria) |
-Macromolecule #1: Mgp-operon protein 3
| Macromolecule | Name: Mgp-operon protein 3 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycoplasmoides genitalium G37 (bacteria) Mycoplasmoides genitalium G37 (bacteria) |
| Molecular weight | Theoretical: 114.553211 KDa |
| Sequence | String: MKTMRKQIYK KAYWLLLPFL PLALANTFLV KEDSKNVTAY TPFATPITDS KSDLVSLAQL DSSYQIADQT IHNTNLFVLF KSRDVKVKY ESSGSNNISF DSTSQGEKPS YVVEFTNSTN IGIKWTMVKK YQLDVPNVSS DMNQVLKNLI LEQPLTKYTL N SSLAKEKG ...String: MKTMRKQIYK KAYWLLLPFL PLALANTFLV KEDSKNVTAY TPFATPITDS KSDLVSLAQL DSSYQIADQT IHNTNLFVLF KSRDVKVKY ESSGSNNISF DSTSQGEKPS YVVEFTNSTN IGIKWTMVKK YQLDVPNVSS DMNQVLKNLI LEQPLTKYTL N SSLAKEKG KTQREVHLGS GQANQWTSQR NQHDLNNNPS PNASTGFKLT TGNAYRKLSE SWPIYEPIDG TKQGKGKDSS GW SSTEENE AKNDAPSVSG GGSSSGTFNK YLNTKQALES IGILFDDQTP RNVITQLYYA STSKLAVTNN HIVVMGNSFL PSM WYWVVE RSAQENASNK PTWFANTNLD WGEDKQKQFV ENQLGYKETT STNSHNFHSK SFTQPAYLIS GIDSVNDQII FSGF KAGSV GYDSSSSSSS SSSSSTKDQA LAWSTTTSLD SKTGYKDLVT NDTGLNGPIN GSFSIQDTFS FVVPYSGNHT NNGTT GPIK TAYPVKKDQK STVKINSLIN ATPLNSYGDE GIGVFDALGL NYNFKSNQER LPSRTDQIFV YGIVSPNELR SAKSSA DST GSDTKVNWSN TQSRYLPVPY NYSEGIIDAD GFKRPENRGA SVTTFSGLKS IAPDGFANSI ANFSVGLKAG IDPNPVM SG KKANYGAVVL TRGGVVRLNF NPGNDSLLST TDNNIAPISF SFTPFTAAES AVDLTTFKEV TYNQESGLWS YIFDSSLK P SHDGKQTPVT DNMGFSVITV SRTGIELNQD QATTTLDVAP SALAVQSGIQ STTQTLTGVL PLSEEFSAVI AKDSDQNKI DIYKNNNGLF EIDTQLSNSV ATNNGGLAPS YTENRVDAWG KVEFADNSVL QARNLVDKTV DEIINTPEIL NSFFRFTPAF EDQKATLVA TKQSDTSLSV SPRIQFLDGN FYDLNSTIAG VPLNIGFPSR VFAGFAALPA WVIPVSVGSS VGILFILLVL G LGIGIPMY RVRKLQDASF VNVFKKVDTL TTAVGSVYKK IITQTGVVKK APSALKAANP SVKKPAAFLK PPVQPPSKPE GE QKAVEVK SEETKS UniProtKB: Adhesin P110 |
-Macromolecule #2: Adhesin P1
| Macromolecule | Name: Adhesin P1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycoplasmoides genitalium G37 (bacteria) Mycoplasmoides genitalium G37 (bacteria) |
| Molecular weight | Theoretical: 159.809297 KDa |
| Sequence | String: MHQPKKRLAK KSWAFLTAAL TLGVITGVGG YFLFNQNKQR SSVSNFAYQP KQLSVKHQQA VDETLTPWTW NNNNFSSLKI TGENPGSFG LVRSQNDNLN ISSVTKNSSD DNLKYLNAVE KYLDGQQNFA IRRYDNNGRA LYDINLAKME NPSTVQRGLN G EPIFDPFK ...String: MHQPKKRLAK KSWAFLTAAL TLGVITGVGG YFLFNQNKQR SSVSNFAYQP KQLSVKHQQA VDETLTPWTW NNNNFSSLKI TGENPGSFG LVRSQNDNLN ISSVTKNSSD DNLKYLNAVE KYLDGQQNFA IRRYDNNGRA LYDINLAKME NPSTVQRGLN G EPIFDPFK GFGLTGNAPT DWNEIKGKVP VEVVQSPHSP NLYFVLLVPK VALEYHNLNN QVVKESLEVK ATQSSFNPTQ RL QKDSPVK DSSKQGEKLS ETTASSMSSG MATSTRAKAL KVEVERGSQS DSLLKNDFAK KPLKHKNSSG EVKLEAEKEF TEA WKPLLT TDQIAREKGM GATVVSFYDA PYSENHTAFG LVDHIDPKKM VENYPPSWKT PKWNHHGIWD YNARNLLLQT TGFF NPRRH PEWFDEGQAK ADNTSPGFKV GDTDHKKDGF KKNSSSPIAL PFEAYFANIG NMVAIGNSVF IFGGNGHATK MFTTN PLSI GVFRIKYTDN FSKSSVTGWP YAVLFGGLIN PQTNGLKDLP LGTNRWFEYV PRMAVSGVKW VGNQLVLAGT LTMGDT ATV PRLKYDQLEK HLNLVAQGQG LLREDLQIFT PYGWANRPDI PVGAWLQDEM GSKFGPHYFL NNPDIQDNVN NDTVEAL IS SYKNTDKLKH VYPYRYSGLY AWQLFNWSNK LTNTPLSANF VNENSYAPNS LFAAILNEDL LTGLSDKIFY GKENEFAE N EADRFNQLLS LNPNPNTNWA RYLNVVQRFT TGPNLDSSTF DQFLDFLPWI GNGKPFSNSP SPSTSASSST PLPTFSNIN VGVKSMITQH LNKENTRWVF IPNFSPDIWT GAGYRVQSAN QKNGIPFEQV KPSNNSTPFD PNSDDNKVTP SGGSSKPTTY PALPNSISP TSDWINALTF TNKNNPQRNQ LLLRSLLGTI PVLINKSGDS NDQFNKDSEQ KWDKTETNEG NLPGFGEVNG L YNAALLHT YGFFGTNTNS TDPKIGFKAD SSSSSSSTLV GSGLNWTSQD VGNLVVINDT SFGFQLGGWF ITFTDFIRPR TG YLGITLS SLQDQTIIWA DQPWTSFKGS YLDSDGTPKS LWDPTALKSL PNSSTTYDTN PTLSPSFQLY QPNKVKAYQT TNT YNKLIE PVDATSAATN MTSLLKLLTT KNIKAKLGKG TASSQGNNNG GGVSQTINTI TTTGNISEGL KEETSIQAET LKKF FDSKQ NNKSEIGIGD STFTKMDGKL TGVVSTPLVN LINGQGATSD SDTEKISFKP GNQIDFNRLF TLPVTELFDP NTMFV YDQY VPLLVNLPSG FDQASIRLKV ISYSVENQTL GVRLEFKDPQ TQQFIPVLNA SSTGPQTVFQ PFNQWADYVL PLIVTV PIV VIILSVTLGL TIGIPMHRNK KALQAGFDLS NKKVDVLTKA VGSVFKEIIN RTGISNAPKK LKQATPTKPT PKTPPKP PV KQ UniProtKB: Adhesin P140 |
-Macromolecule #4: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Macromolecule #5: PHOSPHATE ION
| Macromolecule | Name: PHOSPHATE ION / type: ligand / ID: 5 / Number of copies: 1 / Formula: PO4 |
|---|---|
| Molecular weight | Theoretical: 94.971 Da |
| Chemical component information |  ChemComp-PO4: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 99 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
| Details | Mildy lysed M. genitalium cells |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)