+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tail needle of bacteriophage SU10 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | bacteriophage / tail / tail needle / VIRUS | |||||||||
| Biological species |   Escherichia phage vB_EcoP_SU10 (virus) Escherichia phage vB_EcoP_SU10 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 10.0 Å | |||||||||
 Authors Authors | Siborova M / Fuzik T / Prochazkova M / Novacek J / Plevka P | |||||||||
| Funding support |  Czech Republic, 1 items Czech Republic, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Tail proteins of phage SU10 reorganize into the nozzle for genome delivery. Authors: Marta Šiborová / Tibor Füzik / Michaela Procházková / Jiří Nováček / Martin Benešík / Anders S Nilsson / Pavel Plevka /   Abstract: Escherichia coli phage SU10 belongs to the genus Kuravirus from the class Caudoviricetes of phages with short non-contractile tails. In contrast to other short-tailed phages, the tails of Kuraviruses ...Escherichia coli phage SU10 belongs to the genus Kuravirus from the class Caudoviricetes of phages with short non-contractile tails. In contrast to other short-tailed phages, the tails of Kuraviruses elongate upon cell attachment. Here we show that the virion of SU10 has a prolate head, containing genome and ejection proteins, and a tail, which is formed of portal, adaptor, nozzle, and tail needle proteins and decorated with long and short fibers. The binding of the long tail fibers to the receptors in the outer bacterial membrane induces the straightening of nozzle proteins and rotation of short tail fibers. After the re-arrangement, the nozzle proteins and short tail fibers alternate to form a nozzle that extends the tail by 28 nm. Subsequently, the tail needle detaches from the nozzle proteins and five types of ejection proteins are released from the SU10 head. The nozzle with the putative extension formed by the ejection proteins enables the delivery of the SU10 genome into the bacterial cytoplasm. It is likely that this mechanism of genome delivery, involving the formation of the tail nozzle, is employed by all Kuraviruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14909.map.gz emd_14909.map.gz | 80.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14909-v30.xml emd-14909-v30.xml emd-14909.xml emd-14909.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14909_fsc.xml emd_14909_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_14909.png emd_14909.png | 39.1 KB | ||
| Filedesc metadata |  emd-14909.cif.gz emd-14909.cif.gz | 5.1 KB | ||
| Others |  emd_14909_half_map_1.map.gz emd_14909_half_map_1.map.gz emd_14909_half_map_2.map.gz emd_14909_half_map_2.map.gz | 80.8 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14909 http://ftp.pdbj.org/pub/emdb/structures/EMD-14909 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14909 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14909 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14909.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14909.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.38 Å | ||||||||||||||||||||||||||||||||||||
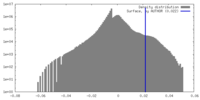
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_14909_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
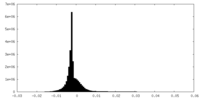
| Density Histograms |
-Half map: #2
| File | emd_14909_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Escherichia phage vB_EcoP_SU10
| Entire | Name:  Escherichia phage vB_EcoP_SU10 (virus) Escherichia phage vB_EcoP_SU10 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Escherichia phage vB_EcoP_SU10
| Supramolecule | Name: Escherichia phage vB_EcoP_SU10 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: NCBI taxonomy ID: 1519788 / NCBI-ID: 1519788 / Sci species name: Escherichia phage vB_EcoP_SU10 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Molecular weight | Theoretical: 105.8 kDa/nm |
| Virus shell | Shell ID: 1 / Name: Tail needle |
-Macromolecule #1: Tail Needle
| Macromolecule | Name: Tail Needle / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MGAGGFRKNT GRNNTTLPYN VGFLNNVQDT ETYNVAVYDE LQKV STATN QMFQAIDEIH EEIDVRIKAL NAMNLQFDEL ENRITTEIET AIADIQTQMG NLS TEDIWD NSVQPPVKLE STVSGFKTSI EGNTTKIQTV EGIVNDQGQE IALVQTELQN IN GSLSQYM ...String: MGAGGFRKNT GRNNTTLPYN VGFLNNVQDT ETYNVAVYDE LQKV STATN QMFQAIDEIH EEIDVRIKAL NAMNLQFDEL ENRITTEIET AIADIQTQMG NLS TEDIWD NSVQPPVKLE STVSGFKTSI EGNTTKIQTV EGIVNDQGQE IALVQTELQN IN GSLSQYM KLSEYEATWG VNSTVNGRYA GVKLTNNGTN SQFQVTANKF IVGDGSSGNT P FVFEGGRA RMEFADIKNV NITTAQIANA RIQWAQIDNV SISNAQIQNL SADKITAGSM WGSNWRLTV GGDFVMGGTG GAQLWMNGNR IDFYDGSGAL RIRIGSW GENBANK: GENBANK: NC_027395.1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: OTHER | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | PFU 10^11 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Average exposure time: 1.0 sec. / Average electron dose: 49.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



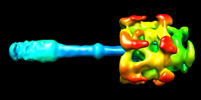





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)