[English] 日本語
 Yorodumi
Yorodumi- EMDB-13454: Active Melanocortin-4 receptor (MC4R)- Gs protein complex bound t... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13454 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Active Melanocortin-4 receptor (MC4R)- Gs protein complex bound to agonist NDP-alpha-MSH at 2.86 A resolution. | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | GPCR / MELANOCORTIN-4 RECEPTOR / MELANOCORTIN RECEPTORS / SETMELANOTIDE / NDP-ALPHA-MSH / ALPHA-18 MSH / ANTAGONISM / AGONISM / APPETITE REGULATION / ANTI-OBESITY TREATMENT / SIGNALING PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of eating behavior / response to melanocyte-stimulating hormone / corticotropin receptor activity / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Prostacyclin signalling through prostacyclin receptor / melanocyte-stimulating hormone receptor activity / G alpha (z) signalling events / Glucagon-type ligand receptors / G beta:gamma signalling through PI3Kgamma ...regulation of eating behavior / response to melanocyte-stimulating hormone / corticotropin receptor activity / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Prostacyclin signalling through prostacyclin receptor / melanocyte-stimulating hormone receptor activity / G alpha (z) signalling events / Glucagon-type ligand receptors / G beta:gamma signalling through PI3Kgamma / G beta:gamma signalling through CDC42 / melanocortin receptor activity / Adrenaline,noradrenaline inhibits insulin secretion / ADP signalling through P2Y purinoceptor 12 / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / negative regulation of eating behavior / Thromboxane signalling through TP receptor / G beta:gamma signalling through BTK / Thrombin signalling through proteinase activated receptors (PARs) / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G alpha (s) signalling events / Ca2+ pathway / G-protein activation / regulation of feeding behavior / Extra-nuclear estrogen signaling / G alpha (12/13) signalling events / G alpha (q) signalling events / Vasopressin regulates renal water homeostasis via Aquaporins / GPER1 signaling / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADP signalling through P2Y purinoceptor 1 / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G-protein activation / Activation of the phototransduction cascade / Glucagon-type ligand receptors / Thromboxane signalling through TP receptor / Sensory perception of sweet, bitter, and umami (glutamate) taste / G beta:gamma signalling through PI3Kgamma / G beta:gamma signalling through CDC42 / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / positive regulation of bone resorption / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / feeding behavior / Ca2+ pathway / G alpha (z) signalling events / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Adrenaline,noradrenaline inhibits insulin secretion / ADP signalling through P2Y purinoceptor 12 / G alpha (q) signalling events / G alpha (i) signalling events / neuropeptide binding / Thrombin signalling through proteinase activated receptors (PARs) / insulin secretion / photoreceptor outer segment membrane / spectrin binding / response to food / alkylglycerophosphoethanolamine phosphodiesterase activity / adenylate cyclase-activating G protein-coupled bile acid receptor signaling pathway / adenylate cyclase-activating serotonin receptor signaling pathway / regulation of skeletal muscle contraction / PKA activation in glucagon signalling / hair follicle placode formation / developmental growth / intracellular transport / photoreceptor outer segment / D1 dopamine receptor binding / vascular endothelial cell response to laminar fluid shear stress / renal water homeostasis / activation of adenylate cyclase activity / Hedgehog 'off' state / cellular response to acidic pH / adenylate cyclase-activating adrenergic receptor signaling pathway / cardiac muscle cell apoptotic process / photoreceptor inner segment / Transcriptional and post-translational regulation of MITF-M expression and activity / cellular response to glucagon stimulus / intracellular glucose homeostasis / Peptide ligand-binding receptors / positive regulation of insulin secretion involved in cellular response to glucose stimulus / adenylate cyclase activator activity / trans-Golgi network membrane / negative regulation of inflammatory response to antigenic stimulus / response to prostaglandin E / response to insulin / bone development / platelet aggregation / cognition / G-protein beta/gamma-subunit complex binding / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / positive regulation of insulin secretion / Glucagon signaling in metabolic regulation / Prostacyclin signalling through prostacyclin receptor / sensory perception of smell / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) / Homo sapiens (human) / synthetic construct (others) /    | ||||||||||||
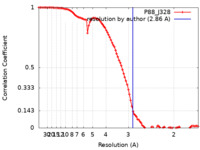
| Method | single particle reconstruction / cryo EM / Resolution: 2.86 Å | ||||||||||||
 Authors Authors | Heyder NA / Schmidt A / Kleinau G / Hilal T / Scheerer P | ||||||||||||
| Funding support |  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell Res / Year: 2021 Journal: Cell Res / Year: 2021Title: Structures of active melanocortin-4 receptor-Gs-protein complexes with NDP-α-MSH and setmelanotide. Authors: Nicolas A Heyder / Gunnar Kleinau / David Speck / Andrea Schmidt / Sarah Paisdzior / Michal Szczepek / Brian Bauer / Anja Koch / Monique Gallandi / Dennis Kwiatkowski / Jörg Bürger / ...Authors: Nicolas A Heyder / Gunnar Kleinau / David Speck / Andrea Schmidt / Sarah Paisdzior / Michal Szczepek / Brian Bauer / Anja Koch / Monique Gallandi / Dennis Kwiatkowski / Jörg Bürger / Thorsten Mielke / Annette G Beck-Sickinger / Peter W Hildebrand / Christian M T Spahn / Daniel Hilger / Magdalena Schacherl / Heike Biebermann / Tarek Hilal / Peter Kühnen / Brian K Kobilka / Patrick Scheerer /   Abstract: The melanocortin-4 receptor (MC4R), a hypothalamic master regulator of energy homeostasis and appetite, is a class A G-protein-coupled receptor and a prime target for the pharmacological treatment of ...The melanocortin-4 receptor (MC4R), a hypothalamic master regulator of energy homeostasis and appetite, is a class A G-protein-coupled receptor and a prime target for the pharmacological treatment of obesity. Here, we present cryo-electron microscopy structures of MC4R-Gs-protein complexes with two drugs recently approved by the FDA, the peptide agonists NDP-α-MSH and setmelanotide, with 2.9 Å and 2.6 Å resolution. Together with signaling data from structure-derived MC4R mutants, the complex structures reveal the agonist-induced origin of transmembrane helix (TM) 6-regulated receptor activation. The ligand-binding modes of NDP-α-MSH, a high-affinity linear variant of the endogenous agonist α-MSH, and setmelanotide, a cyclic anti-obesity drug with biased signaling toward Gq/11, underline the key role of TM3 in ligand-specific interactions and of calcium ion as a ligand-adaptable cofactor. The agonist-specific TM3 interplay subsequently impacts receptor-Gs-protein interfaces at intracellular loop 2, which also regulates the G-protein coupling profile of this promiscuous receptor. Finally, our structures reveal mechanistic details of MC4R activation/inhibition, and provide important insights into the regulation of the receptor signaling profile which will facilitate the development of tailored anti-obesity drugs. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13454.map.gz emd_13454.map.gz | 4.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13454-v30.xml emd-13454-v30.xml emd-13454.xml emd-13454.xml | 23.5 KB 23.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13454_fsc.xml emd_13454_fsc.xml | 9.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_13454.png emd_13454.png | 67 KB | ||
| Filedesc metadata |  emd-13454.cif.gz emd-13454.cif.gz | 7.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13454 http://ftp.pdbj.org/pub/emdb/structures/EMD-13454 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13454 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13454 | HTTPS FTP |
-Related structure data
| Related structure data |  7pivMC  7piuC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13454.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13454.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ternary complex of the NDP-alpha-MSH activated melanocortin 4 rec...
| Entire | Name: Ternary complex of the NDP-alpha-MSH activated melanocortin 4 receptor (MC4R) Gs protein complex stabilized by nanobody 35 |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of the NDP-alpha-MSH activated melanocortin 4 rec...
| Supramolecule | Name: Ternary complex of the NDP-alpha-MSH activated melanocortin 4 receptor (MC4R) Gs protein complex stabilized by nanobody 35 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 144.6 kDa/nm |
-Macromolecule #1: Melanocortin receptor 4
| Macromolecule | Name: Melanocortin receptor 4 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.69852 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DYKDDDDKMV NSTHRGMHTS LHLWNRSSYR LHSNASESLG KGYSDGGCYE QLFVSPEVFV TLGVISLLEN ILVIVAIAKN KNLHSPMYF FICSLAVADM LVSVSNGSET IVITLLNSTD TDAQSFTVNI DNVIDSVICS SLLASICSLL SIAVDRYFTI F YALQYHNI ...String: DYKDDDDKMV NSTHRGMHTS LHLWNRSSYR LHSNASESLG KGYSDGGCYE QLFVSPEVFV TLGVISLLEN ILVIVAIAKN KNLHSPMYF FICSLAVADM LVSVSNGSET IVITLLNSTD TDAQSFTVNI DNVIDSVICS SLLASICSLL SIAVDRYFTI F YALQYHNI MTVKRVGIII SCIWAACTVS GILFIIYSDS SAVIICLITM FFTMLALMAS LYVHMFLMAR LHIKRIAVLP GT GAIRQGA NMKGAITLTI LIGVFVVCWA PFFLHLIFYI SCPQNPYCVC FMSHFNLYLI LIMCNSIIDP LIYALRSQEL RKT FKEIIC CYPLGGLCDL SSRYLEVLFQ UniProtKB: Melanocortin receptor 4 |
-Macromolecule #2: NDP-alpha-MSH (other names Afamelanotide; Scenesse)
| Macromolecule | Name: NDP-alpha-MSH (other names Afamelanotide; Scenesse) / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 1.632863 KDa |
| Sequence | String: (ACE)SYS(NLE)EH(DPN)RW GKPV(NH2) |
-Macromolecule #3: Isoform Gnas-2 of Guanine nucleotide-binding protein G(s) subunit...
| Macromolecule | Name: Isoform Gnas-2 of Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.32616 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKSTIVKQM RILHVNGFNG DSEKATKVQD IKNNLKEAI ETIVAAMSNL VPPVELANPE NQFRVDYILS VMNVPDFDFP PEFYEHAKAL WEDEGVRACY ERSNEYQLID C AQYFLDKI ...String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKSTIVKQM RILHVNGFNG DSEKATKVQD IKNNLKEAI ETIVAAMSNL VPPVELANPE NQFRVDYILS VMNVPDFDFP PEFYEHAKAL WEDEGVRACY ERSNEYQLID C AQYFLDKI DVIKQADYVP SDQDLLRCRV LTSGIFETKF QVDKVNFHMF DVGGQRDERR KWIQCFNDVT AIIFVVASSS YN MVIREDN QTNRLQEALN LFKSIWNNRW LRTISVILFL NKQDLLAEKV LAGKSKIEDY FPEFARYTTP EDATPEPGED PRV TRAKYF IRDEFLRIST ASGDGRHYCY PHFTCAVDTE NIRRVFNDCR DIIQRMHLRQ YELL UniProtKB: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 37.728152 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: GPGSSGSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: GPGSSGSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #5: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #6: Camelid antibody VHH fragment - nanobody 35
| Macromolecule | Name: Camelid antibody VHH fragment - nanobody 35 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.71432 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLQESGGG LVQPGGSLRL SCAASGFTFS NYKMNWVRQA PGKGLEWVSD ISQSGASISY TGSVKGRFTI SRDNAKNTLY LQMNSLKPE DTAVYYCARC PAPFTRDCFD VTSTTYAYRG QGTQVTVSSH HHHHH |
-Macromolecule #7: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 7 / Number of copies: 1 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #8: water
| Macromolecule | Name: water / type: ligand / ID: 8 / Number of copies: 115 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.2 mg/mL | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 5618 / Average exposure time: 40.78 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: OTHER | ||||||
| Output model |  PDB-7piv: |
 Movie
Movie Controller
Controller






























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)