+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11978 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Nanobody E bound to Spike-RBD in a localized reconstruction. | |||||||||
 Map data Map data | Resolve Cryo-EM from the Phenix suite used at half-maps from localised reconstruction in CryoSparc. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | spike glycoprotein / SARS-CoV-2 / nanobody / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
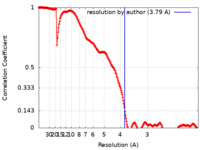
| Method | single particle reconstruction / cryo EM / Resolution: 3.79 Å | |||||||||
 Authors Authors | Hallberg BM / Das H | |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape. Authors: Paul-Albert Koenig / Hrishikesh Das / Hejun Liu / Beate M Kümmerer / Florian N Gohr / Lea-Marie Jenster / Lisa D J Schiffelers / Yonas M Tesfamariam / Miki Uchima / Jennifer D Wuerth / Karl ...Authors: Paul-Albert Koenig / Hrishikesh Das / Hejun Liu / Beate M Kümmerer / Florian N Gohr / Lea-Marie Jenster / Lisa D J Schiffelers / Yonas M Tesfamariam / Miki Uchima / Jennifer D Wuerth / Karl Gatterdam / Natalia Ruetalo / Maria H Christensen / Caroline I Fandrey / Sabine Normann / Jan M P Tödtmann / Steffen Pritzl / Leo Hanke / Jannik Boos / Meng Yuan / Xueyong Zhu / Jonathan L Schmid-Burgk / Hiroki Kato / Michael Schindler / Ian A Wilson / Matthias Geyer / Kerstin U Ludwig / B Martin Hällberg / Nicholas C Wu / Florian I Schmidt /    Abstract: The pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to spread, with devastating consequences. For passive immunization efforts, nanobodies have size and cost ...The pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to spread, with devastating consequences. For passive immunization efforts, nanobodies have size and cost advantages over conventional antibodies. In this study, we generated four neutralizing nanobodies that target the receptor binding domain of the SARS-CoV-2 spike protein. We used x-ray crystallography and cryo-electron microscopy to define two distinct binding epitopes. On the basis of these structures, we engineered multivalent nanobodies with more than 100 times the neutralizing activity of monovalent nanobodies. Biparatopic nanobody fusions suppressed the emergence of escape mutants. Several nanobody constructs neutralized through receptor binding competition, whereas other monovalent and biparatopic nanobodies triggered aberrant activation of the spike fusion machinery. These premature conformational changes in the spike protein forestalled productive fusion and rendered the virions noninfectious. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11978.map.gz emd_11978.map.gz | 751.2 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11978-v30.xml emd-11978-v30.xml emd-11978.xml emd-11978.xml | 25.5 KB 25.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11978_fsc.xml emd_11978_fsc.xml | 20.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_11978.png emd_11978.png | 58 KB | ||
| Filedesc metadata |  emd-11978.cif.gz emd-11978.cif.gz | 7.3 KB | ||
| Others |  emd_11978_half_map_1.map.gz emd_11978_half_map_1.map.gz emd_11978_half_map_2.map.gz emd_11978_half_map_2.map.gz | 763.5 MB 763.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11978 http://ftp.pdbj.org/pub/emdb/structures/EMD-11978 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11978 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11978 | HTTPS FTP |
-Related structure data
| Related structure data |  7b14MC  7b17C  7b18C  7kn5C  7kn6C  7kn7C  7ksgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11978.map.gz / Format: CCP4 / Size: 802.7 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11978.map.gz / Format: CCP4 / Size: 802.7 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Resolve Cryo-EM from the Phenix suite used at half-maps from localised reconstruction in CryoSparc. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.02 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Half map 1 from localised reconstruction
| File | emd_11978_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 from localised reconstruction | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2 from localised reconstruction
| File | emd_11978_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 from localised reconstruction | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 glycoprotein in complex with a nanobody named V
| Entire | Name: SARS-CoV-2 glycoprotein in complex with a nanobody named V |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 glycoprotein in complex with a nanobody named V
| Supramolecule | Name: SARS-CoV-2 glycoprotein in complex with a nanobody named V type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Molecular weight | Theoretical: 510 KDa |
-Supramolecule #2: Spike glycoprotein
| Supramolecule | Name: Spike glycoprotein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Nanobody against SARS-CoV-2
| Supramolecule | Name: Nanobody against SARS-CoV-2 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Spike protein S1
| Macromolecule | Name: Spike protein S1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.002676 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT ...String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT NGVGYQPYRV VVLSFELLHA PATVCGPK UniProtKB: Spike glycoprotein |
-Macromolecule #2: Nanobody against SARS-CoV-2
| Macromolecule | Name: Nanobody against SARS-CoV-2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.109523 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLVETGGG FVQPGGSLRL SCAASGVTLD YYAIGWFRQA PGKEREGVSC IGSSDGRTYY SDSVKGRFTI SRDNAKNTVY LQMNSLKPE DTAVYYCALT VGTYYSGNYH YTCSDDMDYW GKGTQVTVSS |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.3 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 49.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-7b14: |
 Movie
Movie Controller
Controller





















 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)