[English] 日本語
 Yorodumi
Yorodumi- EMDB-11668: 1.15 A structure of human apoferritin obtained from Titan Mono- B... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11668 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 1.15 A structure of human apoferritin obtained from Titan Mono- BCOR microscope | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Apoferritin / METAL BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationiron ion sequestering activity / ferritin complex / Scavenging by Class A Receptors / Golgi Associated Vesicle Biogenesis / ferroxidase / autolysosome / negative regulation of ferroptosis / ferroxidase activity / negative regulation of fibroblast proliferation / ferric iron binding ...iron ion sequestering activity / ferritin complex / Scavenging by Class A Receptors / Golgi Associated Vesicle Biogenesis / ferroxidase / autolysosome / negative regulation of ferroptosis / ferroxidase activity / negative regulation of fibroblast proliferation / ferric iron binding / autophagosome / iron ion transport / ferrous iron binding / Iron uptake and transport / tertiary granule lumen / ficolin-1-rich granule lumen / intracellular iron ion homeostasis / immune response / iron ion binding / negative regulation of cell population proliferation / Neutrophil degranulation / extracellular exosome / extracellular region / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.15 Å | |||||||||
 Authors Authors | Yip KM / Fischer N | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Atomic-resolution protein structure determination by cryo-EM. Authors: Ka Man Yip / Niels Fischer / Elham Paknia / Ashwin Chari / Holger Stark /  Abstract: Single-particle electron cryo-microscopy (cryo-EM) is a powerful method for solving the three-dimensional structures of biological macromolecules. The technological development of transmission ...Single-particle electron cryo-microscopy (cryo-EM) is a powerful method for solving the three-dimensional structures of biological macromolecules. The technological development of transmission electron microscopes, detectors and automated procedures in combination with user-friendly image processing software and ever-increasing computational power have made cryo-EM a successful and expanding technology over the past decade. At resolutions better than 4 Å, atomic model building starts to become possible, but the direct visualization of true atomic positions in protein structure determination requires much higher (better than 1.5 Å) resolution, which so far has not been attained by cryo-EM. The direct visualization of atom positions is essential for understanding the mechanisms of protein-catalysed chemical reactions, and for studying how drugs bind to and interfere with the function of proteins. Here we report a 1.25 Å-resolution structure of apoferritin obtained by cryo-EM with a newly developed electron microscope that provides, to our knowledge, unprecedented structural detail. Our apoferritin structure has almost twice the 3D information content of the current world record reconstruction (at 1.54 Å resolution). We can visualize individual atoms in a protein, see density for hydrogen atoms and image single-atom chemical modifications. Beyond the nominal improvement in resolution, we also achieve a substantial improvement in the quality of the cryo-EM density map, which is highly relevant for using cryo-EM in structure-based drug design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11668.map.gz emd_11668.map.gz | 393.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11668-v30.xml emd-11668-v30.xml emd-11668.xml emd-11668.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11668.png emd_11668.png | 100.6 KB | ||
| Filedesc metadata |  emd-11668.cif.gz emd-11668.cif.gz | 5.9 KB | ||
| Others |  emd_11668_additional.map.gz emd_11668_additional.map.gz emd_11668_additional_1.map.gz emd_11668_additional_1.map.gz | 385.9 MB 385.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11668 http://ftp.pdbj.org/pub/emdb/structures/EMD-11668 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11668 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11668 | HTTPS FTP |
-Related structure data
| Related structure data |  7a6aMC  6z6uC  6z9eC  6z9fC  7a6bC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10591 (Title: Atomic resolution structure of apoferritin from Titan Mono/BCorr microscope EMPIAR-10591 (Title: Atomic resolution structure of apoferritin from Titan Mono/BCorr microscopeData size: 41.7 TB Data #1: Single particle cryo-EM dataset of apoferritin from Titan Mono-BCorr microscope at 1.25 angstrom resolution [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11668.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11668.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.492 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
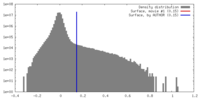
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unsharpened map
| File | emd_11668_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
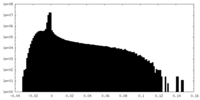
| Density Histograms |
-Additional map: unsharpened map
| File | emd_11668_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Apoferritin
| Entire | Name: Apoferritin |
|---|---|
| Components |
|
-Supramolecule #1: Apoferritin
| Supramolecule | Name: Apoferritin / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Ferritin heavy chain
| Macromolecule | Name: Ferritin heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 24 / Enantiomer: LEVO / EC number: ferroxidase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 21.270605 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTTASTSQVR QNYHQDSEAA INRQINLELY ASYVYLSMSY YFDRDDVALK NFAKYFLHQS HEEREHAEKL MKLQNQRGGR IFLQDIQKP D(CSX)DDWESGLN AMECALHLEK NVNQSLLELH KLATDKNDPH LCDFIETHYL NEQVKAIKEL GDHVTNL RK MGAPESGLAE YLFDKHTLGD SDNES UniProtKB: Ferritin heavy chain |
-Macromolecule #2: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 2 / Number of copies: 32 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 4621 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.5 mg/mL |
|---|---|
| Buffer | pH: 7.6 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: CEOS B-COR / Chromatic aberration corrector: TFS Monochromator |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: O (octahedral) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 1.15 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 797000 |
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)