[English] 日本語
 Yorodumi
Yorodumi- EMDB-11360: Cryo-EM structure of the 90S pre-ribosome from Saccharomyces cere... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11360 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the 90S pre-ribosome from Saccharomyces cerevisiae, state Post-A1 | |||||||||
 Map data Map data | main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosome / 90S pre-ribosome / 40S pre-ribosome / A1 cleavage / Dhr1 | |||||||||
| Function / homology |  Function and homology information Function and homology information18S rRNA (adenine1779-N6/adenine1780-N6)-dimethyltransferase / 18S rRNA (adenine(1779)-N(6)/adenine(1780)-N(6))-dimethyltransferase activity / regulation of ribosomal protein gene transcription by RNA polymerase II / box H/ACA snoRNA binding / RNA fragment catabolic process / rRNA small subunit pseudouridine methyltransferase Nep1 / CURI complex / UTP-C complex / rRNA 2'-O-methylation / Noc4p-Nop14p complex ...18S rRNA (adenine1779-N6/adenine1780-N6)-dimethyltransferase / 18S rRNA (adenine(1779)-N(6)/adenine(1780)-N(6))-dimethyltransferase activity / regulation of ribosomal protein gene transcription by RNA polymerase II / box H/ACA snoRNA binding / RNA fragment catabolic process / rRNA small subunit pseudouridine methyltransferase Nep1 / CURI complex / UTP-C complex / rRNA 2'-O-methylation / Noc4p-Nop14p complex / endonucleolytic cleavage in ITS1 upstream of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / t-UTP complex / Pwp2p-containing subcomplex of 90S preribosome / Mpp10 complex / nuclear microtubule / rRNA (pseudouridine) methyltransferase activity / box C/D sno(s)RNA binding / snoRNA guided rRNA 2'-O-methylation / rRNA modification / histone H2AQ104 methyltransferase activity / septum digestion after cytokinesis / rRNA (adenine-N6,N6-)-dimethyltransferase activity / tRNA re-export from nucleus / snRNA binding / box C/D sno(s)RNA 3'-end processing / rRNA methyltransferase activity / endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / regulation of rRNA processing / regulation of transcription by RNA polymerase I / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / RNA folding chaperone / box C/D methylation guide snoRNP complex / rDNA heterochromatin / U4/U6 snRNP / positive regulation of rRNA processing / tRNA export from nucleus / single-stranded telomeric DNA binding / rRNA primary transcript binding / sno(s)RNA-containing ribonucleoprotein complex / O-methyltransferase activity / : / rRNA base methylation / protein localization to nucleolus / SUMOylation of RNA binding proteins / U4 snRNA binding / rRNA methylation / mTORC1-mediated signalling / Protein hydroxylation / U4 snRNP / 90S preribosome assembly / U3 snoRNA binding / positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / poly(U) RNA binding / Formation of the ternary complex, and subsequently, the 43S complex / Translation initiation complex formation / poly(A)+ mRNA export from nucleus / Ribosomal scanning and start codon recognition / snoRNA binding / preribosome, small subunit precursor / precatalytic spliceosome / Major pathway of rRNA processing in the nucleolus and cytosol / establishment of cell polarity / positive regulation of transcription by RNA polymerase I / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / nucleolar large rRNA transcription by RNA polymerase I / spliceosomal complex assembly / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / L13a-mediated translational silencing of Ceruloplasmin expression / 90S preribosome / Ub-specific processing proteases / RNA processing / ribosomal subunit export from nucleus / proteasome assembly / U4/U6 x U5 tri-snRNP complex / regulation of translational fidelity / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal small subunit export from nucleus / RNA endonuclease activity / nuclear periphery / Transferases; Transferring one-carbon groups; Methyltransferases / helicase activity / ribosome assembly / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / spliceosomal complex / maturation of SSU-rRNA / small-subunit processome / translational initiation / mRNA splicing, via spliceosome / maintenance of translational fidelity / enzyme activator activity / rRNA processing / : / peroxisome / ribosomal small subunit assembly / ribosome biogenesis / ribosomal small subunit biogenesis Similarity search - Function | |||||||||
| Biological species |   | |||||||||
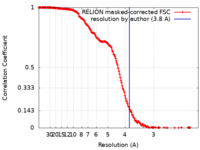
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Cheng J / Lau B | |||||||||
 Citation Citation |  Journal: Science / Year: 2020 Journal: Science / Year: 2020Title: 90 pre-ribosome transformation into the primordial 40 subunit. Authors: Jingdong Cheng / Benjamin Lau / Giuseppe La Venuta / Michael Ameismeier / Otto Berninghausen / Ed Hurt / Roland Beckmann /  Abstract: Production of small ribosomal subunits initially requires the formation of a 90 precursor followed by an enigmatic process of restructuring into the primordial pre-40 subunit. We elucidate this ...Production of small ribosomal subunits initially requires the formation of a 90 precursor followed by an enigmatic process of restructuring into the primordial pre-40 subunit. We elucidate this process by biochemical and cryo-electron microscopy analysis of intermediates along this pathway in yeast. First, the remodeling RNA helicase Dhr1 engages the 90 pre-ribosome, followed by Utp24 endonuclease-driven RNA cleavage at site A, thereby separating the 5'-external transcribed spacer (ETS) from 18 ribosomal RNA. Next, the 5'-ETS and 90 assembly factors become dislodged, but this occurs sequentially, not en bloc. Eventually, the primordial pre-40 emerges, still retaining some 90 factors including Dhr1, now ready to unwind the final small nucleolar U3-18 RNA hybrid. Our data shed light on the elusive 90 to pre-40 transition and clarify the principles of assembly and remodeling of large ribonucleoproteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11360.map.gz emd_11360.map.gz | 52.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11360-v30.xml emd-11360-v30.xml emd-11360.xml emd-11360.xml | 109.6 KB 109.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11360_fsc.xml emd_11360_fsc.xml | 17.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_11360.png emd_11360.png | 187.6 KB | ||
| Filedesc metadata |  emd-11360.cif.gz emd-11360.cif.gz | 28.2 KB | ||
| Others |  emd_11360_additional_1.map.gz emd_11360_additional_1.map.gz emd_11360_additional_2.map.gz emd_11360_additional_2.map.gz emd_11360_additional_3.map.gz emd_11360_additional_3.map.gz emd_11360_additional_4.map.gz emd_11360_additional_4.map.gz | 10 MB 18 MB 9 MB 9.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11360 http://ftp.pdbj.org/pub/emdb/structures/EMD-11360 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11360 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11360 | HTTPS FTP |
-Related structure data
| Related structure data |  6zqdMC  6zqaC  6zqbC  6zqcC  6zqeC  6zqfC  6zqgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11360.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11360.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.084 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: multibody refine focus Head
| File | emd_11360_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | multibody refine focus Head | ||||||||||||
| Projections & Slices |
| ||||||||||||
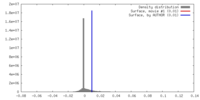
| Density Histograms |
-Additional map: Multibody refine focus Body
| File | emd_11360_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Multibody refine focus Body | ||||||||||||
| Projections & Slices |
| ||||||||||||
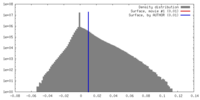
| Density Histograms |
-Additional map: Multibody refine focus Platform
| File | emd_11360_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Multibody refine focus Platform | ||||||||||||
| Projections & Slices |
| ||||||||||||
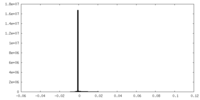
| Density Histograms |
-Additional map: Multibody refine focus Base
| File | emd_11360_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Multibody refine focus Base | ||||||||||||
| Projections & Slices |
| ||||||||||||
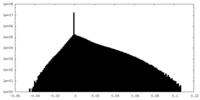
| Density Histograms |
- Sample components
Sample components
+Entire : 90S pre-ribbosome state Post-A1
+Supramolecule #1: 90S pre-ribbosome state Post-A1
+Macromolecule #1: Periodic tryptophan protein 2
+Macromolecule #2: Nucleolar complex protein 14
+Macromolecule #3: Something about silencing protein 10
+Macromolecule #4: U3 small nucleolar RNA-associated protein 4
+Macromolecule #5: U3 small nucleolar RNA-associated protein 5
+Macromolecule #6: U3 small nucleolar RNA-associated protein 6
+Macromolecule #7: U3 small nucleolar RNA-associated protein 7
+Macromolecule #8: U3 small nucleolar RNA-associated protein 8
+Macromolecule #9: U3 small nucleolar RNA-associated protein 9
+Macromolecule #10: U3 small nucleolar RNA-associated protein 10
+Macromolecule #11: U3 small nucleolar RNA-associated protein 11
+Macromolecule #12: U3 small nucleolar RNA-associated protein 12
+Macromolecule #13: U3 small nucleolar RNA-associated protein 13
+Macromolecule #14: U3 small nucleolar RNA-associated protein 14
+Macromolecule #15: U3 small nucleolar RNA-associated protein 15
+Macromolecule #16: Bud site selection protein 21
+Macromolecule #17: NET1-associated nuclear protein 1
+Macromolecule #18: U3 small nucleolar RNA-associated protein 18
+Macromolecule #19: Nucleolar complex protein 4
+Macromolecule #20: U3 small nucleolar RNA-associated protein 20
+Macromolecule #21: U3 small nucleolar RNA-associated protein 21
+Macromolecule #22: U3 small nucleolar RNA-associated protein 22
+Macromolecule #23: rRNA-processing protein FCF1
+Macromolecule #24: rRNA 2'-O-methyltransferase fibrillarin
+Macromolecule #25: Nucleolar protein 56
+Macromolecule #26: Nucleolar protein 58
+Macromolecule #27: 13 kDa ribonucleoprotein-associated protein
+Macromolecule #28: Ribosomal RNA-processing protein 9
+Macromolecule #29: U3 small nucleolar ribonucleoprotein protein IMP3
+Macromolecule #30: U3 small nucleolar ribonucleoprotein protein IMP4
+Macromolecule #31: U3 small nucleolar RNA-associated protein MPP10
+Macromolecule #32: Ribosome biogenesis protein BMS1
+Macromolecule #33: RNA 3'-terminal phosphate cyclase-like protein
+Macromolecule #34: Ribosomal RNA-processing protein 7
+Macromolecule #35: Probable ATP-dependent RNA helicase DHR1
+Macromolecule #36: Ribosomal RNA small subunit methyltransferase NEP1
+Macromolecule #37: Essential nuclear protein 1
+Macromolecule #38: rRNA biogenesis protein RRP5
+Macromolecule #39: Dimethyladenosine transferase
+Macromolecule #40: rRNA-processing protein FCF2
+Macromolecule #41: Protein SOF1
+Macromolecule #42: 40S ribosomal protein S27-A
+Macromolecule #43: Pre-rRNA-processing protein PNO1
+Macromolecule #44: 40S ribosomal protein S1-A
+Macromolecule #45: 40S ribosomal protein S4-A
+Macromolecule #46: Rps5p
+Macromolecule #47: 40S ribosomal protein S6-A
+Macromolecule #48: 40S ribosomal protein S7-A
+Macromolecule #49: 40S ribosomal protein S8-A
+Macromolecule #50: 40S ribosomal protein S9-A
+Macromolecule #51: 40S ribosomal protein S11-A
+Macromolecule #52: 40S ribosomal protein S13
+Macromolecule #53: 40S ribosomal protein S14-A
+Macromolecule #54: 40S ribosomal protein S16-A
+Macromolecule #55: 40S ribosomal protein S18-A
+Macromolecule #56: 40S ribosomal protein S19-A
+Macromolecule #57: 40S ribosomal protein S22-A
+Macromolecule #58: 40S ribosomal protein S23-A
+Macromolecule #59: 40S ribosomal protein S24-A
+Macromolecule #60: 40S ribosomal protein S28-A
+Macromolecule #61: 5ETS RNA
+Macromolecule #62: 18S rRNA
+Macromolecule #63: U3 snoRNA
+Macromolecule #64: ZINC ION
+Macromolecule #65: GUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #66: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil / Material: COPPER / Support film - Material: CARBON |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: GATAN K2 SUMMIT (4k x 4k) / #0 - Average electron dose: 44.0 e/Å2 / #1 - Image recording ID: 2 / #1 - Film or detector model: FEI FALCON II (4k x 4k) / #1 - Average electron dose: 28.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)