[English] 日本語
 Yorodumi
Yorodumi- EMDB-10738: Structure of the native full-length HIV-1 capsid protein in compl... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10738 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the native full-length HIV-1 capsid protein in complex with Cyclophilin A from helical assembly (-8,13) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV / capsid / hexamer / helical assembly / curvature / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationSynthesis And Processing Of GAG, GAGPOL Polyproteins / host cellular component / host cell nuclear membrane / negative regulation of protein K48-linked ubiquitination / regulation of apoptotic signaling pathway / cell adhesion molecule production / lipid droplet organization / negative regulation of viral life cycle / heparan sulfate binding / regulation of viral genome replication ...Synthesis And Processing Of GAG, GAGPOL Polyproteins / host cellular component / host cell nuclear membrane / negative regulation of protein K48-linked ubiquitination / regulation of apoptotic signaling pathway / cell adhesion molecule production / lipid droplet organization / negative regulation of viral life cycle / heparan sulfate binding / regulation of viral genome replication / virion binding / leukocyte chemotaxis / negative regulation of stress-activated MAPK cascade / activation of protein kinase B activity / endothelial cell activation / Integration of viral DNA into host genomic DNA / Autointegration results in viral DNA circles / Basigin interactions / protein peptidyl-prolyl isomerization / cyclosporin A binding / Minus-strand DNA synthesis / Plus-strand DNA synthesis / Uncoating of the HIV Virion / 2-LTR circle formation / Vpr-mediated nuclear import of PICs / Early Phase of HIV Life Cycle / viral budding via host ESCRT complex / Integration of provirus / APOBEC3G mediated resistance to HIV-1 infection / negative regulation of protein phosphorylation / viral release from host cell / Calcineurin activates NFAT / Binding and entry of HIV virion / negative regulation of protein kinase activity / negative regulation of oxidative stress-induced intrinsic apoptotic signaling pathway / positive regulation of viral genome replication / neutrophil chemotaxis / Membrane binding and targetting of GAG proteins / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / : / HIV-1 retropepsin / peptidylprolyl isomerase / positive regulation of protein secretion / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / peptidyl-prolyl cis-trans isomerase activity / exoribonuclease H / exoribonuclease H activity / Assembly Of The HIV Virion / DNA integration / Budding and maturation of HIV virion / platelet activation / viral genome integration into host DNA / positive regulation of protein phosphorylation / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / platelet aggregation / integrin binding / viral penetration into host nucleus / host multivesicular body / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / neuron differentiation / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / SARS-CoV-1 activates/modulates innate immune responses / : / Platelet degranulation / host cell / protein folding / viral nucleocapsid / cellular response to oxidative stress / DNA recombination / secretory granule lumen / DNA-directed DNA polymerase / vesicle / aspartic-type endopeptidase activity / ficolin-1-rich granule lumen / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / positive regulation of MAPK cascade / symbiont-mediated suppression of host gene expression / viral translational frameshifting / focal adhesion / apoptotic process / Neutrophil degranulation / symbiont entry into host cell / lipid binding / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / protein-containing complex / proteolysis / : / DNA binding / RNA binding / extracellular exosome / extracellular region / zinc ion binding Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
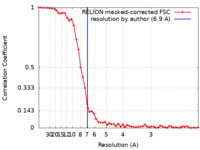
| Method | single particle reconstruction / cryo EM / Resolution: 6.9 Å | |||||||||
 Authors Authors | Ni T / Gerard S | |||||||||
| Funding support |  United Kingdom, United Kingdom,  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2020 Journal: Nat Struct Mol Biol / Year: 2020Title: Intrinsic curvature of the HIV-1 CA hexamer underlies capsid topology and interaction with cyclophilin A. Authors: Tao Ni / Samuel Gerard / Gongpu Zhao / Kyle Dent / Jiying Ning / Jing Zhou / Jiong Shi / Jordan Anderson-Daniels / Wen Li / Sooin Jang / Alan N Engelman / Christopher Aiken / Peijun Zhang /   Abstract: The mature retrovirus capsid consists of a variably curved lattice of capsid protein (CA) hexamers and pentamers. High-resolution structures of the curved assembly, or in complex with host factors, ...The mature retrovirus capsid consists of a variably curved lattice of capsid protein (CA) hexamers and pentamers. High-resolution structures of the curved assembly, or in complex with host factors, have not been available. By devising cryo-EM methodologies for exceedingly flexible and pleomorphic assemblies, we have determined cryo-EM structures of apo-CA hexamers and in complex with cyclophilin A (CypA) at near-atomic resolutions. The CA hexamers are intrinsically curved, flexible and asymmetric, revealing the capsomere and not the previously touted dimer or trimer interfaces as the key contributor to capsid curvature. CypA recognizes specific geometries of the curved lattice, simultaneously interacting with three CA protomers from adjacent hexamers via two noncanonical interfaces, thus stabilizing the capsid. By determining multiple structures from various helical symmetries, we further revealed the essential plasticity of the CA molecule, which allows formation of continuously curved conical capsids and the mechanism of capsid pattern sensing by CypA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10738.map.gz emd_10738.map.gz | 4.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10738-v30.xml emd-10738-v30.xml emd-10738.xml emd-10738.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10738_fsc.xml emd_10738_fsc.xml | 5.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_10738.png emd_10738.png | 79.7 KB | ||
| Masks |  emd_10738_msk_1.map emd_10738_msk_1.map | 11.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10738.cif.gz emd-10738.cif.gz | 6.1 KB | ||
| Others |  emd_10738_half_map_1.map.gz emd_10738_half_map_1.map.gz emd_10738_half_map_2.map.gz emd_10738_half_map_2.map.gz | 10.2 MB 10.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10738 http://ftp.pdbj.org/pub/emdb/structures/EMD-10738 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10738 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10738 | HTTPS FTP |
-Related structure data
| Related structure data |  6y9vMC  6skkC  6skmC  6sknC  6slqC  6sluC  6smuC  6y9wC  6y9xC  6y9yC  6y9zC  6yj5C  6zdjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10738.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10738.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10738_msk_1.map emd_10738_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
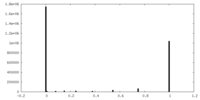
| Density Histograms |
-Half map: #2
| File | emd_10738_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_10738_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : In vitro assembled HIV-1 capsid in tubular assembly
| Entire | Name: In vitro assembled HIV-1 capsid in tubular assembly |
|---|---|
| Components |
|
-Supramolecule #1: In vitro assembled HIV-1 capsid in tubular assembly
| Supramolecule | Name: In vitro assembled HIV-1 capsid in tubular assembly / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: HIV-1 capsid
| Supramolecule | Name: HIV-1 capsid / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Supramolecule #3: Cyclophilin A
| Supramolecule | Name: Cyclophilin A / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Gag-Pol polyprotein
| Macromolecule | Name: Gag-Pol polyprotein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number: HIV-1 retropepsin |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 24.531094 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PIVQNIQGQM VHQAISPRTL NAWVKVVEEK AFSPEVIPMF SALSEGATPQ DLNTMLNTVG GHQAAMQMLK ETINEEAAEW DRVHPVHAG PIAPGQMREP RGSDIAGTTS TLQEQIGWMT NNPPIPVGEI YKRWIILGLN KIVRMYSPTS ILDIRQGPKE P FRDYVDRF ...String: PIVQNIQGQM VHQAISPRTL NAWVKVVEEK AFSPEVIPMF SALSEGATPQ DLNTMLNTVG GHQAAMQMLK ETINEEAAEW DRVHPVHAG PIAPGQMREP RGSDIAGTTS TLQEQIGWMT NNPPIPVGEI YKRWIILGLN KIVRMYSPTS ILDIRQGPKE P FRDYVDRF YKTLRAEQAS QEVKNWMTET LLVQNANPDC KTILKALGPA ATLEEMMTAC QG UniProtKB: Gag-Pol polyprotein |
-Macromolecule #2: Peptidyl-prolyl cis-trans isomerase A
| Macromolecule | Name: Peptidyl-prolyl cis-trans isomerase A / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: peptidylprolyl isomerase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 17.905307 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VNPTVFFDIA VDGEPLGRVS FELFADKVPK TAENFRALST GEKGFGYKGS CFHRIIPGFM CQGGDFTRHN GTGGKSIYGE KFEDENFIL KHTGPGILSM ANAGPNTNGS QFFICTAKTE WLDGKHVVFG KVKEGMNIVE AMERFGSRNG KTSKKITIAD C GQLE UniProtKB: Peptidyl-prolyl cis-trans isomerase A |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R2/1 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE |
| Details | Purified capsid protein were assembled in the presence of Cyclophilin A. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 6500 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 72.14 / Target criteria: Correlation coefficient |
| Output model |  PDB-6y9v: |
 Movie
Movie Controller
Controller


































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)