+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10338 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | XPF-ERCC1 Cryo-EM Structure, DNA-Bound form | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA Repair enzyme. Nucleotide excision repair / DNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of t-circle formation / telomeric DNA-containing double minutes formation / ERCC4-ERCC1 complex / negative regulation of protection from non-homologous end joining at telomere / : / pyrimidine dimer repair by nucleotide-excision repair / nucleotide-excision repair factor 1 complex / nucleotide-excision repair involved in interstrand cross-link repair / nucleotide-excision repair complex / negative regulation of telomere maintenance ...positive regulation of t-circle formation / telomeric DNA-containing double minutes formation / ERCC4-ERCC1 complex / negative regulation of protection from non-homologous end joining at telomere / : / pyrimidine dimer repair by nucleotide-excision repair / nucleotide-excision repair factor 1 complex / nucleotide-excision repair involved in interstrand cross-link repair / nucleotide-excision repair complex / negative regulation of telomere maintenance / single-stranded DNA endonuclease activity / resolution of meiotic recombination intermediates / t-circle formation / mitotic recombination / post-embryonic hemopoiesis / isotype switching / UV protection / negative regulation of telomere maintenance via telomere lengthening / UV-damage excision repair / HDR through Single Strand Annealing (SSA) / oogenesis / TFIID-class transcription factor complex binding / replicative senescence / positive regulation of transcription initiation by RNA polymerase II / response to X-ray / response to UV / interstrand cross-link repair / insulin-like growth factor receptor signaling pathway / telomere maintenance / regulation of autophagy / determination of adult lifespan / nucleotide-excision repair / DNA endonuclease activity / Fanconi Anemia Pathway / promoter-specific chromatin binding / double-strand break repair via homologous recombination / male gonad development / double-strand break repair via nonhomologous end joining / multicellular organism growth / Dual Incision in GG-NER / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / cellular response to UV / single-stranded DNA binding / response to oxidative stress / spermatogenesis / damaged DNA binding / Hydrolases; Acting on ester bonds / protein-macromolecule adaptor activity / chromosome, telomeric region / cell population proliferation / DNA repair / nucleoplasm / identical protein binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.9 Å | |||||||||
 Authors Authors | Jones ML / Briggs DC | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Cryo-EM structures of the XPF-ERCC1 endonuclease reveal how DNA-junction engagement disrupts an auto-inhibited conformation. Authors: Morgan Jones / Fabienne Beuron / Aaron Borg / Andrea Nans / Christopher P Earl / David C Briggs / Ambrosius P Snijders / Maureen Bowles / Edward P Morris / Mark Linch / Neil Q McDonald /  Abstract: The structure-specific endonuclease XPF-ERCC1 participates in multiple DNA damage repair pathways including nucleotide excision repair (NER) and inter-strand crosslink repair (ICLR). How XPF-ERCC1 is ...The structure-specific endonuclease XPF-ERCC1 participates in multiple DNA damage repair pathways including nucleotide excision repair (NER) and inter-strand crosslink repair (ICLR). How XPF-ERCC1 is catalytically activated by DNA junction substrates is not currently understood. Here we report cryo-electron microscopy structures of both DNA-free and DNA-bound human XPF-ERCC1. DNA-free XPF-ERCC1 adopts an auto-inhibited conformation in which the XPF helical domain masks the ERCC1 (HhH) domain and restricts access to the XPF catalytic site. DNA junction engagement releases the ERCC1 (HhH) domain to couple with the XPF-ERCC1 nuclease/nuclease-like domains. Structure-function data indicate xeroderma pigmentosum patient mutations frequently compromise the structural integrity of XPF-ERCC1. Fanconi anaemia patient mutations in XPF often display substantial in-vitro activity but are resistant to activation by ICLR recruitment factor SLX4. Our data provide insights into XPF-ERCC1 architecture and catalytic activation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10338.map.gz emd_10338.map.gz | 20.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10338-v30.xml emd-10338-v30.xml emd-10338.xml emd-10338.xml | 15.6 KB 15.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10338.png emd_10338.png | 57.3 KB | ||
| Filedesc metadata |  emd-10338.cif.gz emd-10338.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10338 http://ftp.pdbj.org/pub/emdb/structures/EMD-10338 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10338 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10338 | HTTPS FTP |
-Related structure data
| Related structure data |  6sxbMC  6sxaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10338.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10338.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.38 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
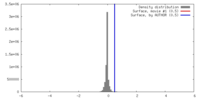
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ERCC1/XPF/DNA complex
| Entire | Name: ERCC1/XPF/DNA complex |
|---|---|
| Components |
|
-Supramolecule #1: ERCC1/XPF/DNA complex
| Supramolecule | Name: ERCC1/XPF/DNA complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: ERCC1/XPF
| Supramolecule | Name: ERCC1/XPF / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: DNA
| Supramolecule | Name: DNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: DNA repair endonuclease XPF
| Macromolecule | Name: DNA repair endonuclease XPF / type: protein_or_peptide / ID: 1 Details: Some residues have been modelled as Alanine where sidechains were not visible. Number of copies: 1 / Enantiomer: LEVO / EC number: Hydrolases; Acting on ester bonds |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 104.636156 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MESGQPARRI AMAPLLEYER QLVLELLDTD GLVVCARGLG ADRLLYHFLQ LHCHPACLVL VLNTQPAEEE YFINQLKIEG VEHLPRRVT NEITSNSRYE VYTQGGVIFA TSRILVVDFL TDRIPSDLIT GILVYRAHRI IESCQEAFIL RLFRQKNKRG F IKAFTDNA ...String: MESGQPARRI AMAPLLEYER QLVLELLDTD GLVVCARGLG ADRLLYHFLQ LHCHPACLVL VLNTQPAEEE YFINQLKIEG VEHLPRRVT NEITSNSRYE VYTQGGVIFA TSRILVVDFL TDRIPSDLIT GILVYRAHRI IESCQEAFIL RLFRQKNKRG F IKAFTDNA VAFDTGFCHV ERVMRNLFVR KLYLWPRFHV AVNSFLEQHK PEVVEIHVSM TPTMLAIQTA ILDILNACLK EL KCHNPSL EVEDLSLENA IGKPFDKTIR HYLDPLWHQL GAKTKSLVQD LKILRTLLQY LSQYDCVTFL NLLESLRATE KAF GQNSGW LFLDSSTSMF INARARVYHL PDAKMSKKEK ISEKMEIKEG EETKKELVLE SNPKWEALTE VLKEIEAENK ESEA LGGPG QVLICASDDR TCSQLRDYIT LGAEAFLLRL YRKTFEKDSK AEEVWMKFRK EDSSKRIRKS HKRPKDPQNK ERAST KERT LKKKKRKLTL TQMVGKPEEL EEEGDVEEGY RREISSSPES CPEEIKHEEF DVNLSSDAAF GILKEPLTII HPLLGC SDP YALTRVLHEV EPRYVVLYDA ELTFVRQLEI YRASRPGKPL RVYFLIYGGS TEEQRYLTAL RKEKEAFEKL IREKASM VV PEEREGRDET NLDLVRGTAS ADVSTDTRKA GGQEQNGTQQ SIVVDMREFR SELPSLIHRR GIDIEPVTLE VGDYILTP E MCVERKSISD LIGSLNNGRL YSQCISMSRY YKRPVLLIEF DPSKPFSLTS RGALFQEISS NDISSKLTLL TLHFPRLRI LWCPSPHATA ELFEELKQSK PQPDAATALA ITADSETLPE SEKYNPGPQD FLLKMPGVNA KNCRSLMHHV KNIAELAALS QDELTSILG NAANAKQLYD FIHTSFAEVV SKGKGKK UniProtKB: DNA repair endonuclease XPF |
-Macromolecule #2: DNA excision repair protein ERCC-1
| Macromolecule | Name: DNA excision repair protein ERCC-1 / type: protein_or_peptide / ID: 2 Details: Some residues modelled as Alanine where no sidechain information available. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.598301 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDPGKDKEGV PQPSGPPARK KFVIPLDEDE VPPGVAKPLF RSTQSLPTVD TSAQAAPQTY AEYAISQPLE GAGATCPTGS EPLAGETPN QALKPGAKSN SIIVSPRQRG NPVLKFVRNV PWEFGDVIPD YVLGQSTCAL FLSLRYHNLH PDYIHGRLQS L GKNFALRV ...String: MDPGKDKEGV PQPSGPPARK KFVIPLDEDE VPPGVAKPLF RSTQSLPTVD TSAQAAPQTY AEYAISQPLE GAGATCPTGS EPLAGETPN QALKPGAKSN SIIVSPRQRG NPVLKFVRNV PWEFGDVIPD YVLGQSTCAL FLSLRYHNLH PDYIHGRLQS L GKNFALRV LLVQVDVKDP QQALKELAKM CILADCTLIL AWSPEEAGRY LETYKAYEQK PADLLMEKLE QDFVSRVTEC LT TVKSVNK TDSQTLLTTF GSLEQLIAAS REDLALCPGL GPQKARRLFD VLHEPFLKVP UniProtKB: DNA excision repair protein ERCC-1 |
-Macromolecule #3: DNA (5'-D(P*CP*AP*GP*AP*TP*GP*CP*TP*GP*A)-3')
| Macromolecule | Name: DNA (5'-D(P*CP*AP*GP*AP*TP*GP*CP*TP*GP*A)-3') / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.06903 KDa |
| Sequence | String: (DC)(DA)(DG)(DA)(DT)(DG)(DC)(DT)(DG)(DA) |
-Macromolecule #4: DNA (5'-D(*TP*CP*AP*GP*CP*AP*TP*CP*TP*G)-3')
| Macromolecule | Name: DNA (5'-D(*TP*CP*AP*GP*CP*AP*TP*CP*TP*G)-3') / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.019992 KDa |
| Sequence | String: (DT)(DC)(DA)(DG)(DC)(DA)(DT)(DC)(DT)(DG) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 Details: 20 mM HEPES pH 7.8, 150 mM NaCl, 1 mM TCEP, 0.01% CHAPS |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 2-17 / Number real images: 15315 / Average electron dose: 63.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 4.5 µm / Calibrated defocus min: 0.9 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.2 µm / Nominal defocus min: 1.2 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 7.9 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2) / Number images used: 198212 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)