[English] 日本語
 Yorodumi
Yorodumi- EMDB-0592: Architectural principles for Hfq/Crc-mediated regulation of gene ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0592 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Architectural principles for Hfq/Crc-mediated regulation of gene expression Hfq-Crc-amiE 2:4:2 complex | |||||||||
 Map data Map data | Locally sharpened Cryo-EM map of a Hfq-Crc-amiE assembly with stoichiometry 2:4:2, respectively. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Hfq / Crc / amiE / Carbon catabolite repression / RNA-protein interaction / RNA-mediated gene regulation / RNA BINDING PROTEIN / RNA BINDING PROTEIN-RNA-HYDROLASE complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of translation, ncRNA-mediated / regulation of carbon utilization / regulation of RNA stability / quorum sensing / double-stranded DNA 3'-5' DNA exonuclease activity / exodeoxyribonuclease III / regulation of translation / DNA repair / regulation of DNA-templated transcription / RNA binding ...regulation of translation, ncRNA-mediated / regulation of carbon utilization / regulation of RNA stability / quorum sensing / double-stranded DNA 3'-5' DNA exonuclease activity / exodeoxyribonuclease III / regulation of translation / DNA repair / regulation of DNA-templated transcription / RNA binding / metal ion binding / cytosol Similarity search - Function | |||||||||
| Biological species |   Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) | |||||||||
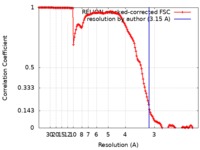
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||
 Authors Authors | Pei XY / Dendooven T | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: Architectural principles for Hfq/Crc-mediated regulation of gene expression. Authors: Xue Yuan Pei / Tom Dendooven / Elisabeth Sonnleitner / Shaoxia Chen / Udo Bläsi / Ben F Luisi /   Abstract: In diverse bacterial species, the global regulator Hfq contributes to post-transcriptional networks that control expression of numerous genes. Hfq of the opportunistic pathogen inhibits translation ...In diverse bacterial species, the global regulator Hfq contributes to post-transcriptional networks that control expression of numerous genes. Hfq of the opportunistic pathogen inhibits translation of target transcripts by forming a regulatory complex with the catabolite repression protein Crc. This repressive complex acts as part of an intricate mechanism of preferred nutrient utilisation. We describe high-resolution cryo-EM structures of the assembly of Hfq and Crc bound to the translation initiation site of a target mRNA. The core of the assembly is formed through interactions of two cognate RNAs, two Hfq hexamers and a Crc pair. Additional Crc protomers are recruited to the core to generate higher-order assemblies with demonstrated regulatory activity in vivo. This study reveals how Hfq cooperates with a partner protein to regulate translation, and provides a structural basis for an RNA code that guides global regulators to interact cooperatively and regulate different RNA targets. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0592.map.gz emd_0592.map.gz | 11.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0592-v30.xml emd-0592-v30.xml emd-0592.xml emd-0592.xml | 23.3 KB 23.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0592_fsc.xml emd_0592_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_0592.png emd_0592.png | 144.8 KB | ||
| Masks |  emd_0592_msk_1.map emd_0592_msk_1.map | 26.2 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-0592.cif.gz emd-0592.cif.gz | 7.1 KB | ||
| Others |  emd_0592_additional.map.gz emd_0592_additional.map.gz emd_0592_half_map_1.map.gz emd_0592_half_map_1.map.gz emd_0592_half_map_2.map.gz emd_0592_half_map_2.map.gz | 24.3 MB 23.3 MB 23.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0592 http://ftp.pdbj.org/pub/emdb/structures/EMD-0592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0592 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0592 | HTTPS FTP |
-Validation report
| Summary document |  emd_0592_validation.pdf.gz emd_0592_validation.pdf.gz | 876 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0592_full_validation.pdf.gz emd_0592_full_validation.pdf.gz | 875.6 KB | Display | |
| Data in XML |  emd_0592_validation.xml.gz emd_0592_validation.xml.gz | 16.9 KB | Display | |
| Data in CIF |  emd_0592_validation.cif.gz emd_0592_validation.cif.gz | 22.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0592 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0592 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0592 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0592 | HTTPS FTP |
-Related structure data
| Related structure data |  6o1mMC  0590C  0591C  6o1kC  6o1lC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0592.map.gz / Format: CCP4 / Size: 26.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0592.map.gz / Format: CCP4 / Size: 26.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally sharpened Cryo-EM map of a Hfq-Crc-amiE assembly with stoichiometry 2:4:2, respectively. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.09 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_0592_msk_1.map emd_0592_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Globally sharpened Cryo-EM map of a Hfq-Crc-amiE assembly...
| File | emd_0592_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Globally sharpened Cryo-EM map of a Hfq-Crc-amiE assembly with stoichiometry 2:4:2, respectively. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_0592_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_0592_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Assembly of Hfq-Crc-amiE with a 2:4:2 stoichiometry
| Entire | Name: Assembly of Hfq-Crc-amiE with a 2:4:2 stoichiometry |
|---|---|
| Components |
|
-Supramolecule #1: Assembly of Hfq-Crc-amiE with a 2:4:2 stoichiometry
| Supramolecule | Name: Assembly of Hfq-Crc-amiE with a 2:4:2 stoichiometry / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 252 KDa |
-Macromolecule #1: Catabolite repression control protein
| Macromolecule | Name: Catabolite repression control protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: exodeoxyribonuclease III |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.101869 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPAMRIISVN VNGIQAAAER GLLSWLQAQN ADVICLQDTR ASAFDLDDPS FQLDGYFLYA CDAELPEQGG VALYSRLQPK AVISGLGFE TADRYGRYLQ ADFDKVSIAT LLLPSGQSGD ESLNQKFKFM DDFTHYLSKQ RRKRREYIYC GSLYVAHQKM D VKNWRECQ ...String: GPAMRIISVN VNGIQAAAER GLLSWLQAQN ADVICLQDTR ASAFDLDDPS FQLDGYFLYA CDAELPEQGG VALYSRLQPK AVISGLGFE TADRYGRYLQ ADFDKVSIAT LLLPSGQSGD ESLNQKFKFM DDFTHYLSKQ RRKRREYIYC GSLYVAHQKM D VKNWRECQ QMPGFLAPER AWLDEVFGNL GYADALREVS REGDQFSWWP DSEQAEMLNL GWRFDYQVLT PGLRRFVRNA KL PRQPRFS QHAPLIVDYD WQLSI UniProtKB: Catabolite repression control protein |
-Macromolecule #2: RNA-binding protein Hfq
| Macromolecule | Name: RNA-binding protein Hfq / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Molecular weight | Theoretical: 7.685984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: HSLQDPYLNT LRKERVPVSI YLVNGIKLQG QIESFDQFVI LLKNTVSQMV YKHAISTVVP SRPVRLP UniProtKB: RNA-binding protein Hfq |
-Macromolecule #3: RNA (5'-R(*AP*AP*AP*AP*AP*UP*AP*AP*CP*AP*AP*CP*AP*AP*GP*AP*GP*G)-3')
| Macromolecule | Name: RNA (5'-R(*AP*AP*AP*AP*AP*UP*AP*AP*CP*AP*AP*CP*AP*AP*GP*AP*GP*G)-3') type: rna / ID: 3 / Number of copies: 2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 5.85766 KDa |
| Sequence | String: AAAAAUAACA ACAAGAGG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Details: unspecified | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K | ||||||||||||
| Details | Monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 70.0 K / Max: 100.0 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 917 / Average exposure time: 60.0 sec. / Average electron dose: 28.0 e/Å2 Details: Images were collected in movie-counting mode, 75 frames per movie stack, 60s exposure per movie |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 130841 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 0.003 µm / Nominal defocus min: 0.00125 µm / Nominal magnification: 125000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross-correlation | ||||||
| Output model |  PDB-6o1m: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)