[English] 日本語
 Yorodumi
Yorodumi- EMDB-0561: Structure of the human frataxin-bound iron-sulfur cluster assembl... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0561 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human frataxin-bound iron-sulfur cluster assembly complex | |||||||||
 Map data Map data | Human frataxin-bound iron-sulfur cluster assembly complex | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
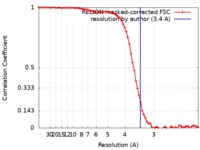
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Fox NG / Yu X | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Structure of the human frataxin-bound iron-sulfur cluster assembly complex provides insight into its activation mechanism. Authors: Nicholas G Fox / Xiaodi Yu / Xidong Feng / Henry J Bailey / Alain Martelli / Joseph F Nabhan / Claire Strain-Damerell / Christine Bulawa / Wyatt W Yue / Seungil Han /   Abstract: The core machinery for de novo biosynthesis of iron-sulfur clusters (ISC), located in the mitochondria matrix, is a five-protein complex containing the cysteine desulfurase NFS1 that is activated by ...The core machinery for de novo biosynthesis of iron-sulfur clusters (ISC), located in the mitochondria matrix, is a five-protein complex containing the cysteine desulfurase NFS1 that is activated by frataxin (FXN), scaffold protein ISCU, accessory protein ISD11, and acyl-carrier protein ACP. Deficiency in FXN leads to the loss-of-function neurodegenerative disorder Friedreich's ataxia (FRDA). Here the 3.2 Å resolution cryo-electron microscopy structure of the FXN-bound active human complex, containing two copies of the NFS1-ISD11-ACP-ISCU-FXN hetero-pentamer, delineates the interactions of FXN with other component proteins of the complex. FXN binds at the interface of two NFS1 and one ISCU subunits, modifying the local environment of a bound zinc ion that would otherwise inhibit NFS1 activity in complexes without FXN. Our structure reveals how FXN facilitates ISC production through stabilizing key loop conformations of NFS1 and ISCU at the protein-protein interfaces, and suggests how FRDA clinical mutations affect complex formation and FXN activation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0561.map.gz emd_0561.map.gz | 3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0561-v30.xml emd-0561-v30.xml emd-0561.xml emd-0561.xml | 14.8 KB 14.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0561_fsc.xml emd_0561_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_0561.png emd_0561.png | 86.8 KB | ||
| Masks |  emd_0561_msk_1.map emd_0561_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Others |  emd_0561_half_map_1.map.gz emd_0561_half_map_1.map.gz emd_0561_half_map_2.map.gz emd_0561_half_map_2.map.gz | 23.4 MB 23.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0561 http://ftp.pdbj.org/pub/emdb/structures/EMD-0561 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0561 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0561 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0561.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0561.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human frataxin-bound iron-sulfur cluster assembly complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.086 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_0561_msk_1.map emd_0561_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
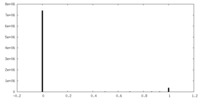
| Density Histograms |
-Half map: Human frataxin-bound iron-sulfur cluster assembly complex, half map 1
| File | emd_0561_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human frataxin-bound iron-sulfur cluster assembly complex, half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
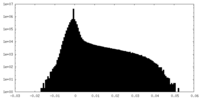
| Density Histograms |
-Half map: Human frataxin-bound iron-sulfur cluster assembly complex, half map 2
| File | emd_0561_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human frataxin-bound iron-sulfur cluster assembly complex, half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
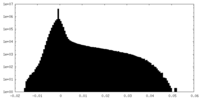
| Density Histograms |
- Sample components
Sample components
-Entire : The human frataxin-bound iron-sulfur cluster assembly complex
| Entire | Name: The human frataxin-bound iron-sulfur cluster assembly complex |
|---|---|
| Components |
|
-Supramolecule #1: The human frataxin-bound iron-sulfur cluster assembly complex
| Supramolecule | Name: The human frataxin-bound iron-sulfur cluster assembly complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Molecular weight | Experimental: 186 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)