+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0458 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
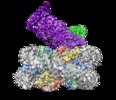
| Title | Structure of Dot1L-H2BK120ub nucleosome complex | |||||||||
 Map data Map data | Structure of Dot1L-H2BK120ub nucleosome complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | chromatin / structural biology / single particle / cryo-EM / histone methyltransferase / nucleosome / Dot1L / GENE REGULATION | |||||||||
| Function / homology |  Function and homology information Function and homology information[histone H3]-lysine79 N-trimethyltransferase / histone H3K79 methyltransferase activity / histone H3K79 trimethyltransferase activity / regulation of transcription regulatory region DNA binding / regulation of receptor signaling pathway via JAK-STAT / histone H3 methyltransferase activity / histone methyltransferase activity / sodium ion transport / subtelomeric heterochromatin formation / telomere organization ...[histone H3]-lysine79 N-trimethyltransferase / histone H3K79 methyltransferase activity / histone H3K79 trimethyltransferase activity / regulation of transcription regulatory region DNA binding / regulation of receptor signaling pathway via JAK-STAT / histone H3 methyltransferase activity / histone methyltransferase activity / sodium ion transport / subtelomeric heterochromatin formation / telomere organization / DNA damage checkpoint signaling / PKMTs methylate histone lysines / structural constituent of chromatin / nucleosome / heterochromatin formation / nucleosome assembly / methylation / gene expression / nucleic acid binding / RNA polymerase II-specific DNA-binding transcription factor binding / mitochondrial outer membrane / transcription coactivator activity / chromosome, telomeric region / protein heterodimerization activity / DNA repair / positive regulation of transcription by RNA polymerase II / protein-containing complex / DNA binding / nucleoplasm / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) / | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Anderson CJ / Baird MR | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2019 Journal: Cell Rep / Year: 2019Title: Structural Basis for Recognition of Ubiquitylated Nucleosome by Dot1L Methyltransferase. Authors: Cathy J Anderson / Matthew R Baird / Allen Hsu / Emily H Barbour / Yuka Koyama / Mario J Borgnia / Robert K McGinty /  Abstract: Histone H3 lysine 79 (H3K79) methylation is enriched on actively transcribed genes, and its misregulation is a hallmark of leukemia. Methylation of H3K79, which resides on the structured disk face of ...Histone H3 lysine 79 (H3K79) methylation is enriched on actively transcribed genes, and its misregulation is a hallmark of leukemia. Methylation of H3K79, which resides on the structured disk face of the nucleosome, is mediated by the Dot1L methyltransferase. Dot1L activity is part of a trans-histone crosstalk pathway, requiring prior histone H2B ubiquitylation of lysine 120 (H2BK120ub) for optimal activity. However, the molecular details describing both how Dot1L binds to the nucleosome and why Dot1L is activated by H2BK120 ubiquitylation are unknown. Here, we present the cryoelectron microscopy (cryo-EM) structure of Dot1L bound to a nucleosome reconstituted with site-specifically ubiquitylated H2BK120. The structure reveals that Dot1L engages the nucleosome acidic patch using a variant arginine anchor and occupies a conformation poised for methylation. In this conformation, Dot1L and ubiquitin interact directly through complementary hydrophobic surfaces. This study establishes a path to better understand Dot1L function in normal and leukemia cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0458.map.gz emd_0458.map.gz | 7.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0458-v30.xml emd-0458-v30.xml emd-0458.xml emd-0458.xml | 25.6 KB 25.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0458_fsc.xml emd_0458_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_0458.png emd_0458.png | 243.3 KB | ||
| Filedesc metadata |  emd-0458.cif.gz emd-0458.cif.gz | 7.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0458 http://ftp.pdbj.org/pub/emdb/structures/EMD-0458 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0458 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0458 | HTTPS FTP |
-Related structure data
| Related structure data |  6nn6MC  0459C  0460C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0458.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0458.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of Dot1L-H2BK120ub nucleosome complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Dot1L-H2BK120ub nucleosome complex (Class 2A)
| Entire | Name: Dot1L-H2BK120ub nucleosome complex (Class 2A) |
|---|---|
| Components |
|
-Supramolecule #1: Dot1L-H2BK120ub nucleosome complex (Class 2A)
| Supramolecule | Name: Dot1L-H2BK120ub nucleosome complex (Class 2A) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#8 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 270 KDa |
-Macromolecule #1: Histone H3.2
| Macromolecule | Name: Histone H3.2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 15.30393 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ARTKQTARKS TGGKAPRKQL ATKAARKSAP ATGGVKKPHR YRPGTVALRE IRRYQKSTEL LIRKLPFQRL VREIAQDFKT DLRFQSSAV MALQEASEAY LVALFEDTNL CAIHAKRVTI MPKDIQLARR IRGERA UniProtKB: Histone H3.2 |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 11.263231 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SGRGKGGKGL GKGGAKRHRK VLRDNIQGIT KPAIRRLARR GGVKRISGLI YEETRGVLKV FLENVIRDAV TYTEHAKRKT VTAMDVVYA LKRQGRTLYG FGG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A type 1
| Macromolecule | Name: Histone H2A type 1 / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 13.978241 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SGRGKQGGKT RAKAKTRSSR AGLQFPVGRV HRLLRKGNYA ERVGAGAPVY LAAVLEYLTA EILELAGNAA RDNKKTRIIP RHLQLAVRN DEELNKLLGR VTIAQGGVLP NIQSVLLPKK TESSKSAKSK UniProtKB: Histone H2A type 1 |
-Macromolecule #4: Histone H2B 1.1
| Macromolecule | Name: Histone H2B 1.1 / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 13.642846 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSAKSAPAPK KGSKKAVTKT QKKDGKKRRK TRKESYAIYV YKVLKQVHPD TGISSKAMSI MNSFVNDVFE RIAGEASRLA HYNKRSTIT SREIQTAVRL LLPGELAKHA VSEGTKAVTC YTSAK UniProtKB: Histone H2B 1.1 |
-Macromolecule #7: Histone-lysine N-methyltransferase, H3 lysine-79 specific
| Macromolecule | Name: Histone-lysine N-methyltransferase, H3 lysine-79 specific type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO / EC number: histone-lysine N-methyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.462039 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSEKLELRLK SPVGAEPAVY PWPLPVYDKH HDAAHEIIET IRWVCEEIPD LKLAMENYVL IDYDTKSFES MQRLCDKYNR AIDSIHQLW KGTTQPMKLN TRPSTGLLRH ILQQVYNHSV TDPEKLNNYE PFSPEVYGET SFDLVAQMID EIKMTDDDLF V DLGSGVGQ ...String: GSEKLELRLK SPVGAEPAVY PWPLPVYDKH HDAAHEIIET IRWVCEEIPD LKLAMENYVL IDYDTKSFES MQRLCDKYNR AIDSIHQLW KGTTQPMKLN TRPSTGLLRH ILQQVYNHSV TDPEKLNNYE PFSPEVYGET SFDLVAQMID EIKMTDDDLF V DLGSGVGQ VVLQVAAATN CKHHYGVEKA DIPAKYAETM DREFRKWMKW YGKKHAEYTL ERGDFLSEEW RERIANTSVI FV NNFAFGP EVDHQLKERF ANMKEGGRIV SSKPFAPLNF RINSRNLSDI GTIMRVVELS PLKGSVSWTG KPVSYYLHTI DRT ILENYF SSLKNPKLRE EQEAARRRQQ RESKSNAATP TKGPEGKVAG PADAPMDSGA EEEKAGAATV KKPSPSKARK KKLN KKGRK MAGRKRGRPK K UniProtKB: Histone-lysine N-methyltransferase, H3 lysine-79 specific |
-Macromolecule #8: Ubiquitin
| Macromolecule | Name: Ubiquitin / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.91118 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSGSMQIFVK TLTGKTITLE VEPSDTIENV KAKIQDKEGI PPDQQRLIFA GKQLEDGRTL SDYNIQKEST LHLVLRLRGC UniProtKB: Ubiquitin C |
-Macromolecule #5: DNA (145-MER)
| Macromolecule | Name: DNA (145-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 44.520383 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC) ...String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC)(DT)(DA) (DG)(DC)(DA)(DC)(DC)(DG)(DC)(DT)(DT)(DA) (DA)(DA) (DC)(DG)(DC)(DA)(DC)(DG)(DT) (DA)(DC)(DG)(DC)(DG)(DC)(DT)(DG)(DT)(DC) (DC)(DC)(DC) (DC)(DG)(DC)(DG)(DT)(DT) (DT)(DT)(DA)(DA)(DC)(DC)(DG)(DC)(DC)(DA) (DA)(DG)(DG)(DG) (DG)(DA)(DT)(DT)(DA) (DC)(DT)(DC)(DC)(DC)(DT)(DA)(DG)(DT)(DC) (DT)(DC)(DC)(DA)(DG) (DG)(DC)(DA)(DC) (DG)(DT)(DG)(DT)(DC)(DA)(DG)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DC)(DA) (DT)(DC)(DG) (DA)(DT) |
-Macromolecule #6: DNA (145-MER)
| Macromolecule | Name: DNA (145-MER) / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 44.99166 KDa |
| Sequence | String: (DA)(DT)(DC)(DG)(DA)(DT)(DG)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA)(DC)(DA) (DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DC)(DT)(DA)(DG)(DG)(DG)(DA) (DG) (DT)(DA)(DA)(DT)(DC)(DC) ...String: (DA)(DT)(DC)(DG)(DA)(DT)(DG)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA)(DC)(DA) (DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DC)(DT)(DA)(DG)(DG)(DG)(DA) (DG) (DT)(DA)(DA)(DT)(DC)(DC)(DC)(DC) (DT)(DT)(DG)(DG)(DC)(DG)(DG)(DT)(DT)(DA) (DA)(DA) (DA)(DC)(DG)(DC)(DG)(DG)(DG) (DG)(DG)(DA)(DC)(DA)(DG)(DC)(DG)(DC)(DG) (DT)(DA)(DC) (DG)(DT)(DG)(DC)(DG)(DT) (DT)(DT)(DA)(DA)(DG)(DC)(DG)(DG)(DT)(DG) (DC)(DT)(DA)(DG) (DA)(DG)(DC)(DT)(DG) (DT)(DC)(DT)(DA)(DC)(DG)(DA)(DC)(DC)(DA) (DA)(DT)(DT)(DG)(DA) (DG)(DC)(DG)(DG) (DC)(DC)(DT)(DC)(DG)(DG)(DC)(DA)(DC)(DC) (DG)(DG)(DG)(DA)(DT)(DT) (DC)(DT)(DG) (DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.93 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 1000 / Average exposure time: 60.0 sec. / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT | ||||||||||||||||||||||||||
| Output model |  PDB-6nn6: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)


























