+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the receptor of xGPR4-Gs complex in pH6.7 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | pH6.7 / xGPR4 / receptor / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcellular response to acidic pH / G protein-coupled receptor activity / adenylate cyclase-activating G protein-coupled receptor signaling pathway / plasma membrane Similarity search - Function | |||||||||
| Biological species | ||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||
 Authors Authors | Rong NK / Wen X / Yang F / Sun JP | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2025 Journal: Cell / Year: 2025Title: Evolutionary study and structural basis of proton sensing by Mus GPR4 and Xenopus GPR4. Authors: Xin Wen / Pan Shang / Haidi Chen / Lulu Guo / Naikang Rong / Xiaoyu Jiang / Xuan Li / Junyan Liu / Gongming Yang / Jiacheng Zhang / Kongkai Zhu / Qingbiao Meng / Xuefei He / Zhihai Wang / ...Authors: Xin Wen / Pan Shang / Haidi Chen / Lulu Guo / Naikang Rong / Xiaoyu Jiang / Xuan Li / Junyan Liu / Gongming Yang / Jiacheng Zhang / Kongkai Zhu / Qingbiao Meng / Xuefei He / Zhihai Wang / Zili Liu / Haoran Cheng / Yilin Zheng / Bifei Zhang / Jiaojiao Pang / Zhaoqian Liu / Peng Xiao / Yuguo Chen / Lunxu Liu / Fengming Luo / Xiao Yu / Fan Yi / Pengju Zhang / Fan Yang / Cheng Deng / Jin-Peng Sun /  Abstract: Animals have evolved pH-sensing membrane receptors, such as G-protein-coupled receptor 4 (GPR4), to monitor pH changes related to their physiology and generate adaptive reactions. However, the ...Animals have evolved pH-sensing membrane receptors, such as G-protein-coupled receptor 4 (GPR4), to monitor pH changes related to their physiology and generate adaptive reactions. However, the evolutionary trajectory and structural mechanism of proton sensing by GPR4 remain unresolved. Here, we observed a positive correlation between the optimal pH of GPR4 activity and the blood pH range across different species. By solving 7-cryoelectron microscopy (cryo-EM) structures of Xenopus tropicalis GPR4 (xtGPR4) and Mus musculus GPR4 (mmGPR4) under varying pH conditions, we identified that protonation of H and H enabled polar network establishment and tighter association between the extracellular loop 2 (ECL2) and 7 transmembrane (7TM) domain, as well as a conserved propagating path, which are common mechanisms underlying protonation-induced GPR4 activation across different species. Moreover, protonation of distinct extracellular H contributed to the more acidic optimal pH range of xtGPR4. Overall, our study revealed common and distinct mechanisms of proton sensing by GPR4, from a structural, functional, and evolutionary perspective. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60052.map.gz emd_60052.map.gz | 95.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60052-v30.xml emd-60052-v30.xml emd-60052.xml emd-60052.xml | 18.6 KB 18.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_60052.png emd_60052.png | 69 KB | ||
| Masks |  emd_60052_msk_1.map emd_60052_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-60052.cif.gz emd-60052.cif.gz | 5.9 KB | ||
| Others |  emd_60052_half_map_1.map.gz emd_60052_half_map_1.map.gz emd_60052_half_map_2.map.gz emd_60052_half_map_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60052 http://ftp.pdbj.org/pub/emdb/structures/EMD-60052 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60052 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60052 | HTTPS FTP |
-Related structure data
| Related structure data |  8zf7MC  8zd1C  8zf4C  8zf6C  8zf9C  8zfaC  8zfbC  8zfcC  8zfdC  8zfeC  9jvgC  9jvhC  9jvmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_60052.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60052.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
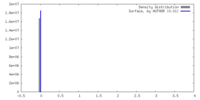
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_60052_msk_1.map emd_60052_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_60052_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_60052_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of the receptor of xGPR4-Gs complex in pH6.7
| Entire | Name: Cryo-EM structure of the receptor of xGPR4-Gs complex in pH6.7 |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of the receptor of xGPR4-Gs complex in pH6.7
| Supramolecule | Name: Cryo-EM structure of the receptor of xGPR4-Gs complex in pH6.7 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: |
-Macromolecule #1: G-protein coupled receptor 4
| Macromolecule | Name: G-protein coupled receptor 4 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 38.083359 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSNFTPDACN VDSGLDSVLP PSLYALVFTL GLPANLLALW AAWLQVRKGR ELGVYLLNLS LSDLLLICAL PPWTDYYLRR DVWGYGPGA CRLFGFVFYT NLYVGAAFLS CVSADRYLAV AHPLRFPGAR PIRSAAAVSA LIWMLELAAN APPLLGEAIH R DRYNHTFC ...String: MSNFTPDACN VDSGLDSVLP PSLYALVFTL GLPANLLALW AAWLQVRKGR ELGVYLLNLS LSDLLLICAL PPWTDYYLRR DVWGYGPGA CRLFGFVFYT NLYVGAAFLS CVSADRYLAV AHPLRFPGAR PIRSAAAVSA LIWMLELAAN APPLLGEAIH R DRYNHTFC YESYPLSGRG AALANVGRVL AGFLLPWGVM MLCYAGLLRA LRGSASCEQR ERRRVRRLAL GLPCVALLCY GP YHALLLL RSLVFLVGGG SVDAGGGCAL EERLFPAYHA SLALATLNCL ADPALYCLAC PGARGEVAKV VGGVVAWAMG KER RAWGER GGNGRGCGEG EEVGMVELRG NGREFVV UniProtKB: G-protein coupled receptor 4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 1.875 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DIFFRACTION / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)