[English] 日本語
 Yorodumi
Yorodumi- EMDB-5017: Segmented eEF2 density from the cryo-EM map of eEF2-bound 80S complex -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5017 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Segmented eEF2 density from the cryo-EM map of eEF2-bound 80S complex | |||||||||
 Map data Map data | Segmented eEF2 density from the 80S.eEF2 cryo-EM map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AlF4- / GDP / GTPase / Ribosome / Translocation | |||||||||
| Function / homology |  Function and homology information Function and homology informationPeptide chain elongation / Synthesis of diphthamide-EEF2 / positive regulation of translational elongation / Protein methylation / protein-synthesizing GTPase / translational elongation / translation elongation factor activity / Neutrophil degranulation / maintenance of translational fidelity / protein-folding chaperone binding ...Peptide chain elongation / Synthesis of diphthamide-EEF2 / positive regulation of translational elongation / Protein methylation / protein-synthesizing GTPase / translational elongation / translation elongation factor activity / Neutrophil degranulation / maintenance of translational fidelity / protein-folding chaperone binding / ribosome binding / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / rRNA binding / ribonucleoprotein complex / GTPase activity / GTP binding / identical protein binding / cytosol Similarity search - Function | |||||||||
| Biological species |   Thermomyces lanuginosus (fungus) Thermomyces lanuginosus (fungus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 12.6 Å | |||||||||
 Authors Authors | Sengupta J / Nilsson J / Gursky R / Kjeldgaard M / Nissen P / Frank J | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2008 Journal: J Mol Biol / Year: 2008Title: Visualization of the eEF2-80S ribosome transition-state complex by cryo-electron microscopy. Authors: Jayati Sengupta / Jakob Nilsson / Richard Gursky / Morten Kjeldgaard / Poul Nissen / Joachim Frank /  Abstract: In an attempt to understand ribosome-induced GTP hydrolysis on eEF2, we determined a 12.6-A cryo-electron microscopy reconstruction of the eEF2-bound 80S ribosome in the presence of aluminum ...In an attempt to understand ribosome-induced GTP hydrolysis on eEF2, we determined a 12.6-A cryo-electron microscopy reconstruction of the eEF2-bound 80S ribosome in the presence of aluminum tetrafluoride and GDP, with aluminum tetrafluoride mimicking the gamma-phosphate during hydrolysis. This is the first visualization of a structure representing a transition-state complex on the ribosome. Tight interactions are observed between the factor's G domain and the large ribosomal subunit, as well as between domain IV and an intersubunit bridge. In contrast, some of the domains of eEF2 implicated in small subunit binding display a large degree of flexibility. Furthermore, we find support for a transition-state model conformation of the switch I region in this complex where the reoriented switch I region interacts with a conserved rRNA region of the 40S subunit formed by loops of the 18S RNA helices 8 and 14. This complex is structurally distinct from the eEF2-bound 80S ribosome complexes previously reported, and analysis of this map sheds light on the GTPase-coupled translocation mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5017.map.gz emd_5017.map.gz | 7.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5017-v30.xml emd-5017-v30.xml emd-5017.xml emd-5017.xml | 9.5 KB 9.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5017_1.png emd_5017_1.png | 105.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5017 http://ftp.pdbj.org/pub/emdb/structures/EMD-5017 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5017 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5017 | HTTPS FTP |
-Related structure data
| Related structure data |  3dnyMC  3dwuMC  5015C  5016C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5017.map.gz / Format: CCP4 / Size: 8.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5017.map.gz / Format: CCP4 / Size: 8.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
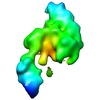
| Annotation | Segmented eEF2 density from the 80S.eEF2 cryo-EM map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.82 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 80S.eEF2.AlF4.GDP
| Entire | Name: 80S.eEF2.AlF4.GDP |
|---|---|
| Components |
|
-Supramolecule #1000: 80S.eEF2.AlF4.GDP
| Supramolecule | Name: 80S.eEF2.AlF4.GDP / type: sample / ID: 1000 Details: GDP-AlF4- traps ribosome-bound eEF2 in a transition state of GTP hydrolysis Number unique components: 4 |
|---|
-Supramolecule #1: 80S ribosome
| Supramolecule | Name: 80S ribosome / type: complex / ID: 1 / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-eukaryote: ALL |
|---|---|
| Source (natural) | Organism:   Thermomyces lanuginosus (fungus) Thermomyces lanuginosus (fungus) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 / Details: 20 mM Hepes-NH3, 100 mM KCl, 20 mM MgCl2 |
|---|---|
| Grid | Details: Quantifoil grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 93 K / Instrument: OTHER / Details: Vitrification instrument: vitrobot |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Category: CCD / Film or detector model: KODAK SO-163 FILM / Digitization - Sampling interval: 2.82 µm / Average electron dose: 10 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Single tilt cryo-holder / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: Segregation in defocus groups and correction in volumes |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 12.6 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Spider / Details: Supervised Classification was used / Number images used: 1 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Details | Protocol: Rigid Body. The domains were separately fitted by manual docking using program O |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: correlation coefficient |
| Output model |  PDB-3dny:  PDB-3dwu: |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















