+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4062 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | MamK double helical filament at 3.6 A resolution | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | bacterial cytoskeleton / filamentous protein / actin-like / magnetosomes / Structural protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationmagnetosome assembly / magnetosome membrane / Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides / cytoskeleton / GTP binding / ATP hydrolysis activity / ATP binding / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Magnetospirillum magneticum AMB-1 (bacteria) Magnetospirillum magneticum AMB-1 (bacteria) | |||||||||
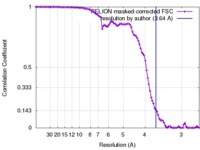
| Method | helical reconstruction / cryo EM / Resolution: 3.64 Å | |||||||||
 Authors Authors | Lowe J | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2016 Journal: Proc Natl Acad Sci U S A / Year: 2016Title: X-ray and cryo-EM structures of monomeric and filamentous actin-like protein MamK reveal changes associated with polymerization. Authors: Jan Löwe / Shaoda He / Sjors H W Scheres / Christos G Savva /  Abstract: Magnetotactic bacteria produce iron-rich magnetic nanoparticles that are enclosed by membrane invaginations to form magnetosomes so they are able to sense and act upon Earth's magnetic field. In ...Magnetotactic bacteria produce iron-rich magnetic nanoparticles that are enclosed by membrane invaginations to form magnetosomes so they are able to sense and act upon Earth's magnetic field. In Magnetospirillum and other magnetotactic bacteria, to combine their magnetic moments, magnetosomes align along filaments formed by a bacterial actin homolog, MamK. Here, we present the crystal structure of a nonpolymerizing mutant of MamK from Magnetospirillum magneticum AMB-1 at 1.8-Å resolution, revealing its close similarity to actin and MreB. The crystals contain AMPPNP-bound monomeric MamK in two different conformations. To investigate conformational changes associated with polymerization, we used unmodified MamK protein and cryo-EM with helical 3D reconstruction in RELION to obtain a density map and a fully refined atomic model of MamK in filamentous form at 3.6-Å resolution. The filament is parallel (polar) double-helical, with a rise of 52.2 Å and a twist of 23.8°. As shown previously and unusually for actin-like filaments, the MamK subunits from each of the two strands are juxtaposed, creating an additional twofold axis along the filament. Compared with monomeric MamK, ADP-bound MamK in the filament undergoes a conformational change, rotating domains I and II against each other to further close the interdomain cleft between subdomains IB and IIB. The domain movement causes several loops to close around the nucleotide-binding pocket. Glu-143, a key residue for catalysis coordinating the magnesium ion, moves closer, presumably switching nucleotide hydrolysis upon polymerization-one of the hallmarks of cytomotive filaments of the actin type. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4062.map.gz emd_4062.map.gz | 78.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4062-v30.xml emd-4062-v30.xml emd-4062.xml emd-4062.xml | 12.5 KB 12.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4062_fsc.xml emd_4062_fsc.xml | 9.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_4062.png emd_4062.png | 167.3 KB | ||
| Filedesc metadata |  emd-4062.cif.gz emd-4062.cif.gz | 5.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4062 http://ftp.pdbj.org/pub/emdb/structures/EMD-4062 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4062 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4062 | HTTPS FTP |
-Related structure data
| Related structure data |  5ljvMC  5ljwC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4062.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4062.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : MamK
| Entire | Name: MamK |
|---|---|
| Components |
|
-Supramolecule #1: MamK
| Supramolecule | Name: MamK / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Magnetospirillum magneticum AMB-1 (bacteria) Magnetospirillum magneticum AMB-1 (bacteria) |
-Macromolecule #1: Actin-like ATPase
| Macromolecule | Name: Actin-like ATPase / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Magnetospirillum magneticum AMB-1 (bacteria) Magnetospirillum magneticum AMB-1 (bacteria) |
| Molecular weight | Theoretical: 37.642152 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSEGEGQAKN RLFLGVDLGT SHTAVMSSRG KKFLLKSVVG YPKDVIGLKL LGRPYVVGDE AFEMRSYLDI RYPLQDGVLS EISDRDIEV ARHLLTHVVK SAEPGPNDEI CAVIGVPARA SAANKALLLK MAQEVVHTAL VVSEPFMVGY GLDKLINTII V DIGAGTTD ...String: MSEGEGQAKN RLFLGVDLGT SHTAVMSSRG KKFLLKSVVG YPKDVIGLKL LGRPYVVGDE AFEMRSYLDI RYPLQDGVLS EISDRDIEV ARHLLTHVVK SAEPGPNDEI CAVIGVPARA SAANKALLLK MAQEVVHTAL VVSEPFMVGY GLDKLINTII V DIGAGTTD ICALKGTVPG PEDQVTLTKA GNYVDERLQN AILERHPELQ MNVNVACAVK EQFSFVGTPT EVASFEFRAA GK PVRADVT EPVKIACEAL MPDIIESIET LLRSFQPEYQ ATVLQNIVFA GGGSRIRGLA AYVKEKLRPF GDANVTCVKD PTF DGCRGA LRLAEELPPQ YWRQLGDVSG S UniProtKB: Actin-like protein MamK |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 6 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil / Material: COPPER/RHODIUM / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4000 pixel / Digitization - Dimensions - Height: 4000 pixel / Number grids imaged: 3 / Number real images: 1665 / Average exposure time: 1.5 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 3.0 µm / Calibrated defocus min: 1.1 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Target criteria: R-factor |
|---|---|
| Output model |  PDB-5ljv: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)