[English] 日本語
 Yorodumi
Yorodumi- EMDB-33560: CryoEM structure of Klebsiella phage Kp9 tail complex applied wit... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of Klebsiella phage Kp9 tail complex applied with C6 symmetry | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / Podophage / short non-contractile tail / VIRUS | |||||||||
| Biological species |  Klebsiella phage Kp9 (virus) Klebsiella phage Kp9 (virus) | |||||||||
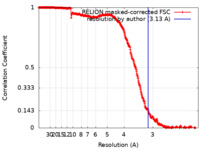
| Method | single particle reconstruction / cryo EM / Resolution: 3.13 Å | |||||||||
 Authors Authors | Huang L / Xiang Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure and assembly of the Klebsiella pneumoniae phage tail fibers Authors: Huang L / Xiang Y | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33560.map.gz emd_33560.map.gz | 128.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33560-v30.xml emd-33560-v30.xml emd-33560.xml emd-33560.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33560_fsc.xml emd_33560_fsc.xml | 35.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_33560.png emd_33560.png | 19.9 KB | ||
| Filedesc metadata |  emd-33560.cif.gz emd-33560.cif.gz | 6.7 KB | ||
| Others |  emd_33560_half_map_1.map.gz emd_33560_half_map_1.map.gz emd_33560_half_map_2.map.gz emd_33560_half_map_2.map.gz | 3 GB 3 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33560 http://ftp.pdbj.org/pub/emdb/structures/EMD-33560 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33560 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33560 | HTTPS FTP |
-Related structure data
| Related structure data |  7y1cMC  7xy1C  7xycC  7y22C  7y23C  7y3tC  7y5sC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33560.map.gz / Format: CCP4 / Size: 3.7 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33560.map.gz / Format: CCP4 / Size: 3.7 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0742 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33560_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33560_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Klebsiella phage Kp9
| Entire | Name:  Klebsiella phage Kp9 (virus) Klebsiella phage Kp9 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Klebsiella phage Kp9
| Supramolecule | Name: Klebsiella phage Kp9 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 2936516 / Sci species name: Klebsiella phage Kp9 / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Klebsiella pneumoniae ATCC 43816 (bacteria) Klebsiella pneumoniae ATCC 43816 (bacteria) |
| Virus shell | Shell ID: 1 / Diameter: 600.0 Å / T number (triangulation number): 7 |
-Macromolecule #1: phage connector protein
| Macromolecule | Name: phage connector protein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Klebsiella phage Kp9 (virus) Klebsiella phage Kp9 (virus) |
| Molecular weight | Theoretical: 58.661219 KDa |
| Sequence | String: MAEVKLEGFA EEGAKAVYDR LKNDRQPYET RAESCAQYTI PSLFPKDSDN ASTDYTTPWQ SVGARGLNNL ASKLMLALFP MQSWMKLTI SEYEAKNLLG DAEGLAKVDE GLSMVERIIM NYIESNSYRV TLFECLKQLC VAGNALLYLP EPEGYTPMKL Y RLNSYVVQ ...String: MAEVKLEGFA EEGAKAVYDR LKNDRQPYET RAESCAQYTI PSLFPKDSDN ASTDYTTPWQ SVGARGLNNL ASKLMLALFP MQSWMKLTI SEYEAKNLLG DAEGLAKVDE GLSMVERIIM NYIESNSYRV TLFECLKQLC VAGNALLYLP EPEGYTPMKL Y RLNSYVVQ RDAFGNVLQI VTLDKIAFNA LPEDVRSQVE AAQGEQKEDA EIDVYTHVYL NEAGDGYSKY EEVAEEVVPG SE AEYPLEE CPYIPVRMVR IDGESYGRSY VEEYLGDLKS LENLQESIVK MAMITAKVIG LVDPAGITQV RRLTAAQSGA FVP GRKQDI EFLQLEKSGD FTVAKNVSDT IEARLSYAFM LNSAVQRTGE RVTAEEIRYV ASELEDTLGG VYSILSQELQ LPLV RVLLK QLQATQQIPE LPKEAVEPTI STGLEAIGRG QDLDKLERCI AAWSALKALE GDDDLNLANL KLRIANAIGL DTAGM LLTQ EQKNALMAQQ GAQIATQQGA AALGQGMAAQ ATASPEAMAA AADSVGMQPG M |
-Macromolecule #2: phage tail tubular protein A
| Macromolecule | Name: phage tail tubular protein A / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Klebsiella phage Kp9 (virus) Klebsiella phage Kp9 (virus) |
| Molecular weight | Theoretical: 21.374646 KDa |
| Sequence | String: MNMQDAYFGS AAELDAVNEM LAAIGESPVT TLDEDGSADV ANARRILNRI NRQIQSKGWA FNINESATLT PDASTGLIPF RPAYLSILG GQYINRGGWV YDKSTGTDTF SGPITVTLIT LQDYDEMPEC FRQWIVTKAS RQFNSRFFGA EDVENSLAQE E MEARMACN ...String: MNMQDAYFGS AAELDAVNEM LAAIGESPVT TLDEDGSADV ANARRILNRI NRQIQSKGWA FNINESATLT PDASTGLIPF RPAYLSILG GQYINRGGWV YDKSTGTDTF SGPITVTLIT LQDYDEMPEC FRQWIVTKAS RQFNSRFFGA EDVENSLAQE E MEARMACN EYEMDFGQYN MLDGDAYVQG LIGR |
-Macromolecule #3: phage tail tubular protein B
| Macromolecule | Name: phage tail tubular protein B / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Klebsiella phage Kp9 (virus) Klebsiella phage Kp9 (virus) |
| Molecular weight | Theoretical: 89.081969 KDa |
| Sequence | String: MALVSQSIKN LKGGISQQPE ILRYPEQGTL QVNGWSSETE GLQKRPPMVF IKSLGPRGYL GEDPYIHLIN RDEYEQYYAV FTGNDVRVF DLSGYEYQVR GDRSYVTVNN PKDNLRMVTV ADYTFIVNRT RQVRENQNRT NGGTFRDNVD AIINVRGGQY G RKLEVNIN ...String: MALVSQSIKN LKGGISQQPE ILRYPEQGTL QVNGWSSETE GLQKRPPMVF IKSLGPRGYL GEDPYIHLIN RDEYEQYYAV FTGNDVRVF DLSGYEYQVR GDRSYVTVNN PKDNLRMVTV ADYTFIVNRT RQVRENQNRT NGGTFRDNVD AIINVRGGQY G RKLEVNIN GVWVSHQLPP GDNAKEDPPK VDAQAIAEAI ATLLRTAHPT WTFNVGTGFI HCIAPADTTI DILETKDGYA DQ LINPVTH YVQSFSKLPL NAPDGYMVKI VGDTSKTADQ YYVKYDKSQK VWKETVGWNI SVGLEYHTMP WTLVRAADGN FDL GYHEWK DRRAGDDDTN PQPSFVNSTI TDVFFFRNRL GFISGENIVM SRTSKYFEFY PPSVANYTDD DPLDVAVSHN RVSV LKYAV SFAEELLLWS DEAQFVLSAN GVLSAKTAQL DLTTQFDVSD RARPYGIGRN IYYASPRSSF TSIMRYYAVQ DVSSV KNAE DMTAHVPNYI PNGVYSINGS GTENFACVLT KGAPSKVFIY KFLYMDENIR QQSWSHWDFG DGVEVMAANC INSTMY MLM RNGYNVWIAA VDFKKESTDF PFEPYRFHVD AKRSYHISET AYDIETNQTV VNVKDIYGAS FAKGTVAICE SDGKITE YE PTGNSWDSTP DIRISGDVSG KNIVIGFLYD FQYVFSRFLI KQEQNDGTTS TEDSGRLQLR RAWVNYQNTG AFTVSVDN G SREFNYLVNA RVGSTGLRLG QKATTTGQYR FPVTGNALYQ KVSLSSFNAS PVSIIGCGWE GNYSRRANGI |
-Macromolecule #4: phage type I tail fiber
| Macromolecule | Name: phage type I tail fiber / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Klebsiella phage Kp9 (virus) Klebsiella phage Kp9 (virus) |
| Molecular weight | Theoretical: 84.120445 KDa |
| Sequence | String: MDQDIKTVIQ YPVGTTEFDI PFDYLSRKFV RVSLVSDDNR RLLSNITEYR YVSKTRVKLL VATTGFDRVE IRRFTSASER IVDFSDGSV LRANDLNVSQ LQSAHIAEEA RDAALLAMPE DDAGNLDARN RKIVRLAPGE AGTDAINKNQ LDTTLGEAGG I LSEVKDLQ ...String: MDQDIKTVIQ YPVGTTEFDI PFDYLSRKFV RVSLVSDDNR RLLSNITEYR YVSKTRVKLL VATTGFDRVE IRRFTSASER IVDFSDGSV LRANDLNVSQ LQSAHIAEEA RDAALLAMPE DDAGNLDARN RKIVRLAPGE AGTDAINKNQ LDTTLGEAGG I LSEVKDLQ KDMEDYLQNW GDDTTAIRGV LWVYNQGSAV GGETSFVITK EGPVLAVPYI EINGSRQYRG WHYEYDLGSK TI TLAKPLS AGDLVVCTTA ETTLPLADSL AGPTGASQIG TANGLNVQIA LDNLRSGVNV LDFMTFAERA AVLNYTGTND NSE AFRKAF ATGSRQIIVP PGRYHVKDVE IPSKVKLFGT YSYKPYNVTS DASFGTDGTI IRKVAGADNM FLWNTACAAE GVMF DGRDR TSPAIQSKSG GKISVGFFKC GFYRFDRVGN RRGAYIGCSF QFCNFNQNNI GIYNTVDGNH IGCTINANKS HGVML ETGA NSNTFTNCRN EWNEGDNWNF YGATSIQVIN ELCDRAFGYG FRISNSSVTL INVNIRRSAR TAASGAASAQ IYFESS TLK MIGVNSSVGG DDTGGSITEP SPDYFFRMAG TSEGRLEISD SRLTGYTVGL ISGTARPSVI RVINSPGWED TINEGVA RI SGGRPYIGTM PTATGPANVS PAVLGLSCGG VNTYDNDMFD IHLTIRNTNN GGHNGAILTV LLYREGGAAR ATIVRVDS R SNAVGEGDVN STSADPQQVY QVSVEVTSND ASTFNLLVST KSDNSASYRF RAKVKP |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 63 |
|---|---|
| Output model |  PDB-7y1c: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)