[English] 日本語
 Yorodumi
Yorodumi- EMDB-31422: Cryo-EM structure of the chemokine receptor CCR5 in complex with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31422 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the chemokine receptor CCR5 in complex with MIP-1a and Gi | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationchemokine (C-C motif) ligand 5 binding / granulocyte chemotaxis / lymphocyte chemotaxis / CCR1 chemokine receptor binding / positive regulation of microglial cell migration / positive regulation of natural killer cell chemotaxis / negative regulation of macrophage apoptotic process / regulation of behavior / signaling / chemokine receptor activity ...chemokine (C-C motif) ligand 5 binding / granulocyte chemotaxis / lymphocyte chemotaxis / CCR1 chemokine receptor binding / positive regulation of microglial cell migration / positive regulation of natural killer cell chemotaxis / negative regulation of macrophage apoptotic process / regulation of behavior / signaling / chemokine receptor activity / astrocyte cell migration / CCR5 chemokine receptor binding / eosinophil degranulation / regulation of sensory perception of pain / negative regulation of bone mineralization / CCR chemokine receptor binding / positive regulation of microglial cell activation / C-C chemokine receptor activity / cell activation / C-C chemokine binding / phosphatidylinositol phospholipase C activity / T cell chemotaxis / positive regulation of calcium ion transport / eosinophil chemotaxis / response to cholesterol / chemokine-mediated signaling pathway / Chemokine receptors bind chemokines / chemokine activity / release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / dendritic cell chemotaxis / phospholipase activator activity / macrophage chemotaxis / positive regulation of calcium ion import / chemoattractant activity / monocyte chemotaxis / negative regulation of osteoclast differentiation / exocytosis / Interleukin-10 signaling / cellular response to interleukin-1 / negative regulation by host of viral transcription / Binding and entry of HIV virion / Adenylate cyclase inhibitory pathway / positive regulation of protein localization to cell cortex / cellular defense response / regulation of cAMP-mediated signaling / coreceptor activity / D2 dopamine receptor binding / G protein-coupled serotonin receptor binding / regulation of mitotic spindle organization / cytoskeleton organization / cellular response to forskolin / neutrophil chemotaxis / positive regulation of calcium-mediated signaling / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / cell chemotaxis / positive regulation of interleukin-1 beta production / Regulation of insulin secretion / G protein-coupled receptor binding / calcium-mediated signaling / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Glucagon signaling in metabolic regulation / G protein-coupled acetylcholine receptor signaling pathway / G-protein activation / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / cellular response to type II interferon / G beta:gamma signalling through CDC42 / response to peptide hormone / Vasopressin regulates renal water homeostasis via Aquaporins / G beta:gamma signalling through BTK / ADP signalling through P2Y purinoceptor 12 / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / Glucagon-type ligand receptors / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / G alpha (z) signalling events / Adrenaline,noradrenaline inhibits insulin secretion / response to toxic substance / intracellular calcium ion homeostasis / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / positive regulation of inflammatory response / osteoblast differentiation / cellular response to catecholamine stimulus / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / adenylate cyclase-activating dopamine receptor signaling pathway / G beta:gamma signalling through PI3Kgamma / GPER1 signaling / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / positive regulation of tumor necrosis factor production / chemotaxis Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Zhang H / Chen K / Tan Q / Han S / Zhu Y / Zhao Q / Wu B | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Structural basis for chemokine recognition and receptor activation of chemokine receptor CCR5. Authors: Hui Zhang / Kun Chen / Qiuxiang Tan / Qiang Shao / Shuo Han / Chenhui Zhang / Cuiying Yi / Xiaojing Chu / Ya Zhu / Yechun Xu / Qiang Zhao / Beili Wu /  Abstract: The chemokine receptor CCR5 plays a vital role in immune surveillance and inflammation. However, molecular details that govern its endogenous chemokine recognition and receptor activation remain ...The chemokine receptor CCR5 plays a vital role in immune surveillance and inflammation. However, molecular details that govern its endogenous chemokine recognition and receptor activation remain elusive. Here we report three cryo-electron microscopy structures of G protein-coupled CCR5 in a ligand-free state and in complex with the chemokine MIP-1α or RANTES, as well as the crystal structure of MIP-1α-bound CCR5. These structures reveal distinct binding modes of the two chemokines and a specific accommodate pattern of the chemokine for the distal N terminus of CCR5. Together with functional data, the structures demonstrate that chemokine-induced rearrangement of toggle switch and plasticity of the receptor extracellular region are critical for receptor activation, while a conserved tryptophan residue in helix II acts as a trigger of receptor constitutive activation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31422.map.gz emd_31422.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31422-v30.xml emd-31422-v30.xml emd-31422.xml emd-31422.xml | 13.7 KB 13.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31422.png emd_31422.png | 26 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31422 http://ftp.pdbj.org/pub/emdb/structures/EMD-31422 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31422 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31422 | HTTPS FTP |
-Validation report
| Summary document |  emd_31422_validation.pdf.gz emd_31422_validation.pdf.gz | 471.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_31422_full_validation.pdf.gz emd_31422_full_validation.pdf.gz | 471.3 KB | Display | |
| Data in XML |  emd_31422_validation.xml.gz emd_31422_validation.xml.gz | 6 KB | Display | |
| Data in CIF |  emd_31422_validation.cif.gz emd_31422_validation.cif.gz | 6.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31422 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31422 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31422 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31422 | HTTPS FTP |
-Related structure data
| Related structure data |  7f1qMC  7f1rC  7f1sC  7f1tC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31422.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31422.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
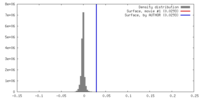
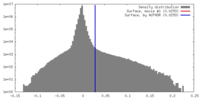
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Chemokine receptor CCR5 in complex with MIP-1a and Gi
| Entire | Name: Chemokine receptor CCR5 in complex with MIP-1a and Gi |
|---|---|
| Components |
|
-Supramolecule #1: Chemokine receptor CCR5 in complex with MIP-1a and Gi
| Supramolecule | Name: Chemokine receptor CCR5 in complex with MIP-1a and Gi / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Macromolecule #1: C-C motif chemokine 3,C-C chemokine receptor type 5
| Macromolecule | Name: C-C motif chemokine 3,C-C chemokine receptor type 5 / type: protein_or_peptide / ID: 1 / Details: Fusion protein of MIP-1a and CCR5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 51.241047 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SLAADTPTAC CFSYCSRQIP QNFIADYFET SSQCSKPGVI FLTKRSRQVC ADPSEEWVQK YVSDLELSAG SGSGSGSGSG SGSGSGSGS GSGSDYQVSS PIYDINYYCS EPCQKINVKQ IAARLLPPLY SLVFIFGFVG NMLVILILIN CKRLKSMTDI Y LLNLAISD ...String: SLAADTPTAC CFSYCSRQIP QNFIADYFET SSQCSKPGVI FLTKRSRQVC ADPSEEWVQK YVSDLELSAG SGSGSGSGSG SGSGSGSGS GSGSDYQVSS PIYDINYYCS EPCQKINVKQ IAARLLPPLY SLVFIFGFVG NMLVILILIN CKRLKSMTDI Y LLNLAISD LFFLLTVPFW AHYAAAQWDF GNTMCQLLTG LYFIGFFSGI FFIILLTIDR YLAVVHAVFA LKARTVTFGV VT SVITWVV AVFASLPNII FTRSQKEGLH YTCSSHFPYS QYQFWKNFQT LKIVILGLVL PLLVMVICYS GILKTLLRCR NEK KRHRAV RLIFTIMIVY FLFWAPYNIV LLLNTFQEFF GLNNCSSSNR LDQAMQVTET LGMTHCCINP IIYAFVGEKF RNYL LVFFQ KHIAKRLEVL FQGPGSWSHP QFEKGSGAGA SAGSWSHPQF EKGSDYKDDD DK |
-Macromolecule #2: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.447141 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKCTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKCTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVTAQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHASM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCS TDTKNVQFVF DAVTDVIIKN NLKDCGLF |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.41693 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL ...String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD INAICFFPNG NA FATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGV TDDGMA VATGSWDSFL KIWN |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 2.1875 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 7095732 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller
















































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)