+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30835 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo EM map of the LAT1-4F2hc bound with JX-075 | ||||||||||||
 Map data Map data | cryo EM map of the LAT1-4F2hc bound with JX-075 | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationL-tryptophan transmembrane transporter activity / alanine transport / L-tryptophan transmembrane transport / positive regulation of L-leucine import across plasma membrane / cellular response to L-arginine / thyroid hormone transmembrane transporter activity / Defective SLC7A7 causes lysinuric protein intolerance (LPI) / neutral L-amino acid secondary active transmembrane transporter activity / apical pole of neuron / amino acid import across plasma membrane ...L-tryptophan transmembrane transporter activity / alanine transport / L-tryptophan transmembrane transport / positive regulation of L-leucine import across plasma membrane / cellular response to L-arginine / thyroid hormone transmembrane transporter activity / Defective SLC7A7 causes lysinuric protein intolerance (LPI) / neutral L-amino acid secondary active transmembrane transporter activity / apical pole of neuron / amino acid import across plasma membrane / aromatic amino acid transmembrane transporter activity / tyrosine transport / L-histidine transport / amino acid transport complex / L-leucine import across plasma membrane / L-alanine transmembrane transporter activity / L-alanine import across plasma membrane / isoleucine transport / phenylalanine transport / methionine transport / L-amino acid transmembrane transporter activity / valine transport / L-leucine transmembrane transporter activity / calcium:sodium antiporter activity / L-leucine transport / thyroid hormone transport / proline transport / positive regulation of cytokine production involved in immune response / amino acid transmembrane transport / negative regulation of vascular associated smooth muscle cell apoptotic process / neutral amino acid transport / amino acid transmembrane transporter activity / external side of apical plasma membrane / Amino acid transport across the plasma membrane / neutral L-amino acid transmembrane transporter activity / Tryptophan catabolism / exogenous protein binding / anchoring junction / response to muscle activity / antiporter activity / Basigin interactions / xenobiotic transport / microvillus membrane / amino acid transport / positive regulation of interleukin-4 production / response to exogenous dsRNA / positive regulation of interleukin-17 production / tryptophan transport / transport across blood-brain barrier / response to hyperoxia / cellular response to glucose starvation / positive regulation of glial cell proliferation / negative regulation of autophagy / basal plasma membrane / liver regeneration / positive regulation of type II interferon production / peptide antigen binding / calcium ion transport / double-stranded RNA binding / melanosome / virus receptor activity / cellular response to lipopolysaccharide / basolateral plasma membrane / carbohydrate metabolic process / cadherin binding / symbiont entry into host cell / protein heterodimerization activity / apical plasma membrane / lysosomal membrane / negative regulation of gene expression / intracellular membrane-bounded organelle / synapse / cell surface / protein homodimerization activity / RNA binding / extracellular exosome / nucleoplasm / membrane / plasma membrane / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | ||||||||||||
 Authors Authors | Yan RH / Li YN / Zhang YY / Zhong XY / Zhou Q | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2021 Journal: Cell Discov / Year: 2021Title: Mechanism of substrate transport and inhibition of the human LAT1-4F2hc amino acid transporter. Authors: Renhong Yan / Yaning Li / Jennifer Müller / Yuanyuan Zhang / Simon Singer / Lu Xia / Xinyue Zhong / Jürg Gertsch / Karl-Heinz Altmann / Qiang Zhou /   Abstract: LAT1 (SLC7A5) is one of the representative light chain proteins of heteromeric amino acid transporters, forming a heterodimer with its heavy chain partner 4F2hc (SLC3A2). LAT1 is overexpressed in ...LAT1 (SLC7A5) is one of the representative light chain proteins of heteromeric amino acid transporters, forming a heterodimer with its heavy chain partner 4F2hc (SLC3A2). LAT1 is overexpressed in many types of tumors and mediates the transfer of drugs and hormones across the blood-brain barrier. Thus, LAT1 is considered as a drug target for cancer treatment and may be exploited for drug delivery into the brain. Here, we synthesized three potent inhibitors of human LAT1, which inhibit transport of leucine with IC values between 100 and 250 nM, and solved the cryo-EM structures of the corresponding LAT1-4F2hc complexes with these inhibitors bound at resolution of up to 2.7 or 2.8 Å. The protein assumes an outward-facing occluded conformation, with the inhibitors bound in the classical substrate binding pocket, but with their tails wedged between the substrate binding site and TM10 of LAT1. We also solved the complex structure of LAT1-4F2hc with 3,5-diiodo-L-tyrosine (Diiodo-Tyr) at 3.4 Å overall resolution, which revealed a different inhibition mechanism and might represent an intermediate conformation between the outward-facing occluded state mentioned above and the outward-open state. To our knowledge, this is the first time that the outward-facing conformation is revealed for the HAT family. Our results unveil more important insights into the working mechanisms of HATs and provide a structural basis for future drug design. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30835.map.gz emd_30835.map.gz | 59.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30835-v30.xml emd-30835-v30.xml emd-30835.xml emd-30835.xml | 13.7 KB 13.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30835.png emd_30835.png | 27.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30835 http://ftp.pdbj.org/pub/emdb/structures/EMD-30835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30835 | HTTPS FTP |
-Validation report
| Summary document |  emd_30835_validation.pdf.gz emd_30835_validation.pdf.gz | 462.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_30835_full_validation.pdf.gz emd_30835_full_validation.pdf.gz | 461.8 KB | Display | |
| Data in XML |  emd_30835_validation.xml.gz emd_30835_validation.xml.gz | 6.1 KB | Display | |
| Data in CIF |  emd_30835_validation.cif.gz emd_30835_validation.cif.gz | 6.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30835 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30835 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30835 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30835 | HTTPS FTP |
-Related structure data
| Related structure data |  7dskMC  7dslC  7dsnC  7dsqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30835.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30835.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo EM map of the LAT1-4F2hc bound with JX-075 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : LAT1-4F2hc bound with JX-075
| Entire | Name: LAT1-4F2hc bound with JX-075 |
|---|---|
| Components |
|
-Supramolecule #1: LAT1-4F2hc bound with JX-075
| Supramolecule | Name: LAT1-4F2hc bound with JX-075 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: 4F2 cell-surface antigen heavy chain
| Macromolecule | Name: 4F2 cell-surface antigen heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 70.160828 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAHHHHHHHH HHSGRELQPP EASIAVVSIP RQLPGSHSEA GVQGLSAGDD SETGSDCVTQ AGLQLLASSD PPALASKNAE VTVETGFHH VSQADIEFLT SIDPTASASG SAGITGTMSQ DTEVDMKEVE LNELEPEKQP MNAASGAAMS LAGAEKNGLV K IKVAEDEA ...String: MAHHHHHHHH HHSGRELQPP EASIAVVSIP RQLPGSHSEA GVQGLSAGDD SETGSDCVTQ AGLQLLASSD PPALASKNAE VTVETGFHH VSQADIEFLT SIDPTASASG SAGITGTMSQ DTEVDMKEVE LNELEPEKQP MNAASGAAMS LAGAEKNGLV K IKVAEDEA EAAAAAKFTG LSKEELLKVA GSPGWVRTRW ALLLLFWLGW LGMLAGAVVI IVRAPRCREL PAQKWWHTGA LY RIGDLQA FQGHGAGNLA GLKGRLDYLS SLKVKGLVLG PIHKNQKDDV AQTDLLQIDP NFGSKEDFDS LLQSAKKKSI RVI LDLTPN YRGENSWFST QVDTVATKVK DALEFWLQAG VDGFQVRDIE NLKDASSFLA EWQNITKGFS EDRLLIAGTN SSDL QQILS LLESNKDLLL TSSYLSDSGS TGEHTKSLVT QYLNATGNRW CSWSLSQARL LTSFLPAQLL RLYQLMLFTL PGTPV FSYG DEIGLDAAAL PGQPMEAPVM LWDESSFPDI PGAVSANMTV KGQSEDPGSL LSLFRRLSDQ RSKERSLLHG DFHAFS AGP GLFSYIRHWD QNERFLVVLN FGDVGLSAGL QASDLPASAS LPAKADLLLS TQPGREEGSP LELERLKLEP HEGLLLR FP YAALE |
-Macromolecule #2: Large neutral amino acids transporter small subunit 1
| Macromolecule | Name: Large neutral amino acids transporter small subunit 1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 57.186895 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADYKDDDDK SGPDEVDASG RAGAGPKRRA LAAPAAEEKE EAREKMLAAK SADGSAPAGE GEGVTLQRNI TLLNGVAIIV GTIIGSGIF VTPTGVLKEA GSPGLALVVW AACGVFSIVG ALCYAELGTT ISKSGGDYAY MLEVYGSLPA FLKLWIELLI I RPSSQYIV ...String: MADYKDDDDK SGPDEVDASG RAGAGPKRRA LAAPAAEEKE EAREKMLAAK SADGSAPAGE GEGVTLQRNI TLLNGVAIIV GTIIGSGIF VTPTGVLKEA GSPGLALVVW AACGVFSIVG ALCYAELGTT ISKSGGDYAY MLEVYGSLPA FLKLWIELLI I RPSSQYIV ALVFATYLLK PLFPTCPVPE EAAKLVACLC VLLLTAVNCY SVKAATRVQD AFAAAKLLAL ALIILLGFVQ IG KGDVSNL DPNFSFEGTK LDVGNIVLAL YSGLFAYGGW NYLNFVTEEM INPYRNLPLA IIISLPIVTL VYVLTNLAYF TTL STEQML SSEAVAVDFG NYHLGVMSWI IPVFVGLSCF GSVNGSLFTS SRLFFVGSRE GHLPSILSMI HPQLLTPVPS LVFT CVMTL LYAFSKDIFS VINFFSFFNW LCVALAIIGM IWLRHRKPEL ERPIKVNLAL PVFFILACLF LIAVSFWKTP VECGI GFTI ILSGLPVYFF GVWWKNKPKW LLQGIFSTTV LCQKLMQVVP QET |
-Macromolecule #4: 1,2-DIACYL-GLYCEROL-3-SN-PHOSPHATE
| Macromolecule | Name: 1,2-DIACYL-GLYCEROL-3-SN-PHOSPHATE / type: ligand / ID: 4 / Number of copies: 1 / Formula: 3PH |
|---|---|
| Molecular weight | Theoretical: 704.998 Da |
| Chemical component information |  ChemComp-3PH: |
-Macromolecule #5: (2~{S})-2-azanyl-7-(naphthalen-1-ylmethoxy)-3,4-dihydro-1~{H}-nap...
| Macromolecule | Name: (2~{S})-2-azanyl-7-(naphthalen-1-ylmethoxy)-3,4-dihydro-1~{H}-naphthalene-2-carboxylic acid type: ligand / ID: 5 / Number of copies: 1 / Formula: HNX |
|---|---|
| Molecular weight | Theoretical: 347.407 Da |
| Chemical component information | 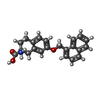 ChemComp-HNX: |
-Macromolecule #6: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 6 / Number of copies: 2 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #7: water
| Macromolecule | Name: water / type: ligand / ID: 7 / Number of copies: 12 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.0.6) / Number images used: 378156 |
|---|---|
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















