+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2646 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the mammalian ribosome-Sec61 complex | |||||||||
 Map data Map data | Mammalian ribosome in complex with Sec61. Class with eEF2 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | translation / ribosome / mammalian / sec61 | |||||||||
| Function / homology |  Function and homology information Function and homology information: / TNFR1-mediated ceramide production / Translation initiation complex formation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / Protein hydroxylation / Degradation of CDH1 / Regulation of TNFR1 signaling / TNFR1-induced NF-kappa-B signaling pathway / multicellular organism development ...: / TNFR1-mediated ceramide production / Translation initiation complex formation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / Protein hydroxylation / Degradation of CDH1 / Regulation of TNFR1 signaling / TNFR1-induced NF-kappa-B signaling pathway / multicellular organism development / L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Major pathway of rRNA processing in the nucleolus and cytosol / Formation of a pool of free 40S subunits / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / translation at presynapse / cellular response to actinomycin D / regulation of G1 to G0 transition / protein-DNA complex disassembly / positive regulation of gastrulation / protein tyrosine kinase inhibitor activity / IRE1-RACK1-PP2A complex / positive regulation of Golgi to plasma membrane protein transport / alpha-beta T cell differentiation / G1 to G0 transition / cysteine-type endopeptidase activator activity involved in apoptotic process / regulation of establishment of cell polarity / negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide / negative regulation of phagocytosis / cytoplasmic side of rough endoplasmic reticulum membrane / organelle membrane / laminin receptor activity / positive regulation of mitochondrial depolarization / negative regulation of Wnt signaling pathway / BH3 domain binding / negative regulation of translational frameshifting / regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway / positive regulation of GTPase activity / regulation of cell division / protein serine/threonine kinase inhibitor activity / ubiquitin ligase inhibitor activity / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / positive regulation of signal transduction by p53 class mediator / protein localization to nucleus / negative regulation of ubiquitin-dependent protein catabolic process / phagocytic cup / positive regulation of intrinsic apoptotic signaling pathway / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / translation regulator activity / rough endoplasmic reticulum / laminin binding / gastrulation / signaling adaptor activity / negative regulation of proteasomal ubiquitin-dependent protein catabolic process / cytosolic ribosome / rescue of stalled cytosolic ribosome / SH2 domain binding / class I DNA-(apurinic or apyrimidinic site) endonuclease activity / cyclin binding / DNA-(apurinic or apyrimidinic site) lyase / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / protein kinase C binding / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / positive regulation of translation / positive regulation of apoptotic signaling pathway / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / negative regulation of smoothened signaling pathway / maturation of SSU-rRNA / small-subunit processome / cellular response to glucose stimulus / positive regulation of protein-containing complex assembly / negative regulation of cell growth / positive regulation of protein phosphorylation / receptor tyrosine kinase binding / maintenance of translational fidelity / modification-dependent protein catabolic process / cellular response to growth factor stimulus / spindle / transcription coactivator binding / protein tag activity / cytoplasmic ribonucleoprotein granule / rRNA processing / rhythmic process / regulation of translation / positive regulation of proteasomal ubiquitin-dependent protein catabolic process / regulation of protein localization / heparin binding / large ribosomal subunit / ribosomal small subunit assembly / ribosome biogenesis / presynapse / virus receptor activity / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / cytosolic small ribosomal subunit Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Voorhees RM / Fernandez IS / Scheres SHW / Hegde R | |||||||||
 Citation Citation |  Journal: Cell / Year: 2014 Journal: Cell / Year: 2014Title: Structure of the mammalian ribosome-Sec61 complex to 3.4 Å resolution. Authors: Rebecca M Voorhees / Israel S Fernández / Sjors H W Scheres / Ramanujan S Hegde /  Abstract: Cotranslational protein translocation is a universally conserved process for secretory and membrane protein biosynthesis. Nascent polypeptides emerging from a translating ribosome are either ...Cotranslational protein translocation is a universally conserved process for secretory and membrane protein biosynthesis. Nascent polypeptides emerging from a translating ribosome are either transported across or inserted into the membrane via the ribosome-bound Sec61 channel. Here, we report structures of a mammalian ribosome-Sec61 complex in both idle and translating states, determined to 3.4 and 3.9 Å resolution. The data sets permit building of a near-complete atomic model of the mammalian ribosome, visualization of A/P and P/E hybrid-state tRNAs, and analysis of a nascent polypeptide in the exit tunnel. Unprecedented chemical detail is observed for both the ribosome-Sec61 interaction and the conformational state of Sec61 upon ribosome binding. Comparison of the maps from idle and translating complexes suggests how conformational changes to the Sec61 channel could facilitate translocation of a secreted polypeptide. The high-resolution structure of the mammalian ribosome-Sec61 complex provides a valuable reference for future functional and structural studies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2646.map.gz emd_2646.map.gz | 31.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2646-v30.xml emd-2646-v30.xml emd-2646.xml emd-2646.xml | 8.5 KB 8.5 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2646.png EMD-2646.png | 330.4 KB | ||
| Others |  emd_2646_half_map_1.map emd_2646_half_map_1.map emd_2646_half_map_2.map emd_2646_half_map_2.map | 282.6 MB 282.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2646 http://ftp.pdbj.org/pub/emdb/structures/EMD-2646 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2646 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2646 | HTTPS FTP |
-Related structure data
| Related structure data |  3j7pMC  2644C  2649C  2650C  3j7oC  3j7qC  3j7rC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2646.map.gz / Format: CCP4 / Size: 276 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2646.map.gz / Format: CCP4 / Size: 276 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mammalian ribosome in complex with Sec61. Class with eEF2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
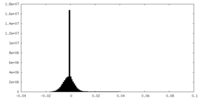
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 2646 half map 1.map
| File | emd_2646_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
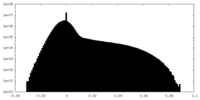
| Density Histograms |
-Supplemental map: emd 2646 half map 2.map
| File | emd_2646_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
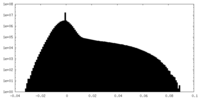
| Density Histograms |
- Sample components
Sample components
-Entire : Mammalian ribosome in complex with Sec61 with the large subunit (...
| Entire | Name: Mammalian ribosome in complex with Sec61 with the large subunit (60S) masked during processing. |
|---|---|
| Components |
|
-Supramolecule #1000: Mammalian ribosome in complex with Sec61 with the large subunit (...
| Supramolecule | Name: Mammalian ribosome in complex with Sec61 with the large subunit (60S) masked during processing. type: sample / ID: 1000 / Number unique components: 2 |
|---|
-Supramolecule #1: Mammalian ribosome
| Supramolecule | Name: Mammalian ribosome / type: complex / ID: 1 / Recombinant expression: No / Ribosome-details: ribosome-eukaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Sec61
| Macromolecule | Name: Sec61 / type: protein_or_peptide / ID: 1 / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 50mM hepes, 200mM K-acetate, 15mM Mg-acetate, 1mM DTT |
|---|---|
| Grid | Details: Quantifoil R2/2 400 mesh copper grids. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 120 K / Instrument: FEI VITROBOT MARK IV Method: 3uL of sampled was incubated on the grid for 30 seconds before blotting for 9 second |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Date | Apr 7, 2014 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Number real images: 1900 / Average electron dose: 25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.003 µm / Nominal defocus min: 0.001 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: OTHER / Software - Name: Relion / Number images used: 80019 |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)