機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / /  データを開く
データを開く 基本情報
基本情報 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報
 データ登録者
データ登録者 米国, 1件
米国, 1件  引用
引用 ジャーナル: Proc Natl Acad Sci U S A / 年: 2021
ジャーナル: Proc Natl Acad Sci U S A / 年: 2021
 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_23008.map.gz
emd_23008.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-23008-v30.xml
emd-23008-v30.xml emd-23008.xml
emd-23008.xml EMDBヘッダ
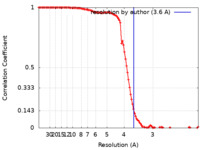
EMDBヘッダ emd_23008_fsc.xml
emd_23008_fsc.xml FSCデータファイル
FSCデータファイル emd_23008.png
emd_23008.png emd-23008.cif.gz
emd-23008.cif.gz emd_23008_additional_1.map.gz
emd_23008_additional_1.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-23008
http://ftp.pdbj.org/pub/emdb/structures/EMD-23008 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23008
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23008 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_23008.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_23008.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)