+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human GBP1 dimer bound to GDP-AlF3 | |||||||||
 Map data Map data | Primary map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cell-autonomous immunity / intracellular pathogens / GTPase / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationGDP phosphatase activity / non-canonical inflammasome complex assembly / negative regulation of substrate adhesion-dependent cell spreading / protein localization to vacuole / positive regulation of pyroptotic inflammatory response / cytolysis in another organism / symbiont cell surface / vesicle membrane / negative regulation of interleukin-2 production / spectrin binding ...GDP phosphatase activity / non-canonical inflammasome complex assembly / negative regulation of substrate adhesion-dependent cell spreading / protein localization to vacuole / positive regulation of pyroptotic inflammatory response / cytolysis in another organism / symbiont cell surface / vesicle membrane / negative regulation of interleukin-2 production / spectrin binding / defense response to protozoan / cytokine binding / cellular response to interleukin-1 / negative regulation of T cell receptor signaling pathway / negative regulation of protein localization to plasma membrane / Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides / regulation of protein localization to plasma membrane / regulation of calcium-mediated signaling / cytoplasmic vesicle membrane / lipopolysaccharide binding / Hsp90 protein binding / negative regulation of ERK1 and ERK2 cascade / cellular response to type II interferon / cellular response to tumor necrosis factor / Interferon gamma signaling / GDP binding / actin cytoskeleton / actin binding / G protein activity / cytoplasmic vesicle / defense response to virus / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / defense response to bacterium / Golgi membrane / innate immune response / GTPase activity / GTP binding / enzyme binding / Golgi apparatus / protein homodimerization activity / extracellular region / identical protein binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
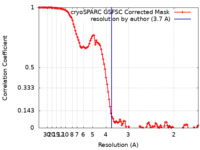
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Kuhm TI / Jakobi AJ | |||||||||
| Funding support | European Union,  Netherlands, 2 items Netherlands, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2025 Journal: Nat Struct Mol Biol / Year: 2025Title: Structural basis of antimicrobial membrane coat assembly by human GBP1. Authors: Tanja Kuhm / Clémence Taisne / Cecilia de Agrela Pinto / Luca Gross / Evdokia A Giannopoulou / Stefan T Huber / Els Pardon / Jan Steyaert / Sander J Tans / Arjen J Jakobi /   Abstract: Guanylate-binding proteins (GBPs) are interferon-inducible guanosine triphosphate hydrolases (GTPases) mediating host defense against intracellular pathogens. Their antimicrobial activity hinges on ...Guanylate-binding proteins (GBPs) are interferon-inducible guanosine triphosphate hydrolases (GTPases) mediating host defense against intracellular pathogens. Their antimicrobial activity hinges on their ability to self-associate and coat pathogen-associated compartments or cytosolic bacteria. Coat formation depends on GTPase activity but how nucleotide binding and hydrolysis prime coat formation remains unclear. Here, we report the cryo-electron microscopy structure of the full-length human GBP1 dimer in its guanine nucleotide-bound state and describe the molecular ultrastructure of the GBP1 coat on liposomes and bacterial lipopolysaccharide membranes. Conformational changes of the middle and GTPase effector domains expose the isoprenylated C terminus for membrane association. The α-helical middle domains form a parallel, crossover arrangement essential for coat formation and position the extended effector domain for intercalation into the lipopolysaccharide layer of gram-negative membranes. Nucleotide binding and hydrolysis create oligomeric scaffolds with contractile abilities that promote membrane extrusion and fragmentation. Our data offer a structural and mechanistic framework for understanding GBP1 effector functions in intracellular immunity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16794.map.gz emd_16794.map.gz | 127.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16794-v30.xml emd-16794-v30.xml emd-16794.xml emd-16794.xml | 24.2 KB 24.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16794_fsc.xml emd_16794_fsc.xml | 10.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_16794.png emd_16794.png | 100.9 KB | ||
| Masks |  emd_16794_msk_1.map emd_16794_msk_1.map | 134.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16794.cif.gz emd-16794.cif.gz | 7.8 KB | ||
| Others |  emd_16794_additional_1.map.gz emd_16794_additional_1.map.gz emd_16794_half_map_1.map.gz emd_16794_half_map_1.map.gz emd_16794_half_map_2.map.gz emd_16794_half_map_2.map.gz | 1.6 MB 124.9 MB 124.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16794 http://ftp.pdbj.org/pub/emdb/structures/EMD-16794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16794 | HTTPS FTP |
-Related structure data
| Related structure data |  8cqbMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16794.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16794.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Primary map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.834 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16794_msk_1.map emd_16794_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Locally sharpened map (LocScale)
| File | emd_16794_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally sharpened map (LocScale) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_16794_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_16794_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Homodimeric complex of GBP1 stabilised by GDP-AlF3
| Entire | Name: Homodimeric complex of GBP1 stabilised by GDP-AlF3 |
|---|---|
| Components |
|
-Supramolecule #1: Homodimeric complex of GBP1 stabilised by GDP-AlF3
| Supramolecule | Name: Homodimeric complex of GBP1 stabilised by GDP-AlF3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 134 KDa |
-Macromolecule #1: Guanylate-binding protein 1
| Macromolecule | Name: Guanylate-binding protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 68.021617 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASEIHMTGP MCLIENTNGR LMANPEALKI LSAITQPMVV VAIVGLYRTG KSYLMNKLAG KKKGFSLGST VQSHTKGIWM WCVPHPKKP GHILVLLDTE GLGDVEKGDN QNDSWIFALA VLLSSTFVYN SIGTINQQAM DQLYYVTELT HRIRSKSSPD E NENEVEDS ...String: MASEIHMTGP MCLIENTNGR LMANPEALKI LSAITQPMVV VAIVGLYRTG KSYLMNKLAG KKKGFSLGST VQSHTKGIWM WCVPHPKKP GHILVLLDTE GLGDVEKGDN QNDSWIFALA VLLSSTFVYN SIGTINQQAM DQLYYVTELT HRIRSKSSPD E NENEVEDS ADFVSFFPDF VWTLRDFSLD LEADGQPLTP DEYLTYSLKL KKGTSQKDET FNLPRLCIRK FFPKKKCFVF DR PVHRRKL AQLEKLQDEE LDPEFVQQVA DFCSYIFSNS KTKTLSGGIQ VNGPRLESLV LTYVNAISSG DLPCMENAVL ALA QIENSA AVQKAIAHYE QQMGQKVQLP TETLQELLDL HRDSEREAIE VFIRSSFKDV DHLFQKELAA QLEKKRDDFC KQNQ EASSD RCSALLQVIF SPLEEEVKAG IYSKPGGYRL FVQKLQDLKK KYYEEPRKGI QAEEILQTYL KSKESMTDAI LQTDQ TLTE KEKEIEVERV KAESAQASAK MLQEMQRKNE QMMEQKERSY QEHLKQLTEK MENDRVQLLK EQERTLALKL QEQEQL LKE GFQKESRIMK NEIQDLQTKM RRRKACTIS UniProtKB: Guanylate-binding protein 1 |
-Macromolecule #2: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 2 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
-Macromolecule #3: ALUMINUM FLUORIDE
| Macromolecule | Name: ALUMINUM FLUORIDE / type: ligand / ID: 3 / Number of copies: 2 / Formula: AF3 |
|---|---|
| Molecular weight | Theoretical: 83.977 Da |
| Chemical component information |  ChemComp-AF3: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: 5u mM HEPES pH7.4, 150 mM NaCl, 1 mM DTT, 200 uM GDP, 10 mM NaF, 300 uM AlCl3, 5 mM MgCl2 | ||||||||||||||||||||||||
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 99 % / Chamber temperature: 22 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 2 / Number real images: 5214 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Target criteria: Real-space cross correlation | ||||||||
| Output model |  PDB-8cqb: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)